













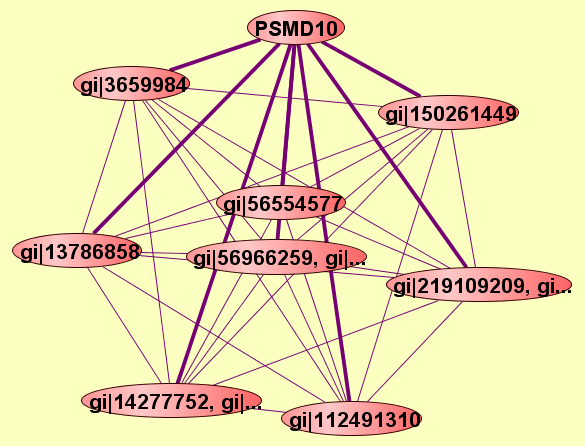
| LEGEND: |
|||
| = Same process, or one is unknown. | = Same branch, distance 4. | ||
| = Same branch, distance 1. | = Same branch, distance 5. | ||
| = Same branch, distance 2. | = Same branch, distance 6 or more. | ||
| = Same branch, distance 3. | = Not in same branch of GO. | ||
| Size | Notes | Members | |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1BD2D and 1UOHA with confidence: 0.927947689664 | PSMD10 gi|3659984 |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 2HE4A and 1UOHA with confidence: 0.98799057509 | PSMD10 gi|112491310 |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 2Q3HA and 1UOHA with confidence: 0.95440856611 | PSMD10 gi|150261449 |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1I1RB and 1UOHA with confidence: 0.956602537135 | PSMD10 gi|13786858 |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1XJDA and 1UOHA with confidence: 0.955485813414 | PSMD10 gi|56554577 |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1I1JA and 1UOHA with confidence: 0.920088516829 | PSMD10 gi|14277752, gi|... |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1UNLA and 1UOHA with confidence: 0.928805833258 | PSMD10 gi|56966260, gi|... |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 2ZHNA and 1UOHA with confidence: 0.980001581023 | PSMD10 gi|219109210, gi... |