













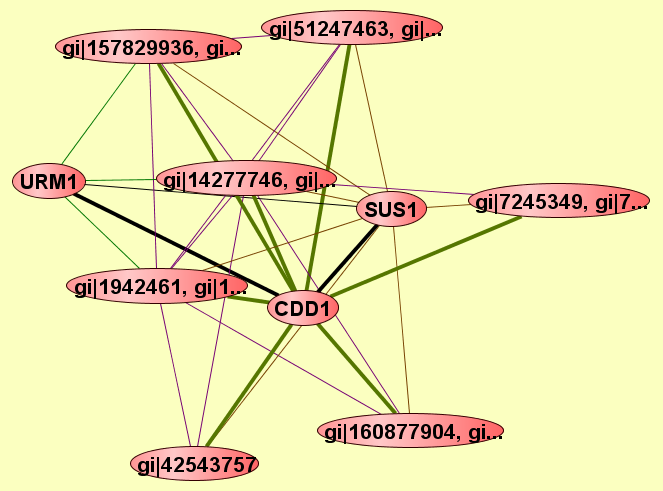
| LEGEND: |
|||
| = Same process, or one is unknown. | = Same branch, distance 4. | ||
| = Same branch, distance 1. | = Same branch, distance 5. | ||
| = Same branch, distance 2. | = Same branch, distance 6 or more. | ||
| = Same branch, distance 3. | = Not in same branch of GO. | ||
| Size | Notes | Members | |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1IGNA and 1R5TA with confidence: 0.98736192687 | CDD1 gi|1942461, gi|1... |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1R5TA and 1D1QA with confidence: 0.901059783997 | CDD1 gi|7245349, gi|7... |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1R5TA and 1I12A with confidence: 0.987578456351 | CDD1 gi|14277743, gi|... |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1R5TA and 1AKYA with confidence: 0.980670051737 | CDD1 gi|157829936, gi... |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1R5TA and 1SZ9A with confidence: 0.904155396485 | CDD1 gi|51247462, gi|... |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1R5TA and 2QJLA with confidence: 0.975871628891 | CDD1 URM1 |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1R5TA and 3FWBC with confidence: 0.989995527752 | CDD1 SUS1 |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1R5TA and 3B5NC with confidence: 0.905045215517 | CDD1 gi|160877908, gi... |
| View GO Analysis | 2 proteins | Predicted to be cocomplexed based on PDB structures 1R5TA and 1USUB with confidence: 0.903648198801 | CDD1 gi|42543757 |