













| General Information: |
|
| Name(s) found: |
SETD3
[HGNC (HUGO)]
|
| Description(s) found:
Found 37 descriptions. SHOW ALL |
|
| Organism: | Homo sapiens |
| Length: | 594 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MGKKSRVKTQ KSGTGATATV SPKEILNLTS ELLQKCSSPA PGPGKEWEEY VQIRTLVEKI 60
61 RKKQKGLSVT FDGKREDYFP DLMKWASENG ASVEGFEMVN FKEEGFGLRA TRDIKAEELF 120
121 LWVPRKLLMT VESAKNSVLG PLYSQDRILQ AMGNIALAFH LLCERASPNS FWQPYIQTLP 180
181 SEYDTPLYFE EDEVRYLQST QAIHDVFSQY KNTARQYAYF YKVIQTHPHA NKLPLKDSFT 240
241 YEDYRWAVSS VMTRQNQIPT EDGSRVTLAL IPLWDMCNHT NGLITTGYNL EDDRCECVAL 300
301 QDFRAGEQIY IFYGTRSNAE FVIHSGFFFD NNSHDRVKIK LGVSKSDRLY AMKAEVLARA 360
361 GIPTSSVFAL HFTEPPISAQ LLAFLRVFCM TEEELKEHLL GDSAIDRIFT LGNSEFPVSW 420
421 DNEVKLWTFL EDRASLLLKT YKTTIEEDKS VLKNHDLSVR AKMAIKLRLG EKEILEKAVK 480
481 SAAVNREYYR QQMEEKAPLP KYEESNLGLL ESSVGDSRLP LVLRNLEEEA GVQDALNIRE 540
541 AISKAKATEN GLVNGENSIP NGTRSENESL NQESKRAVED AKGSSSDSTA GVKE |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
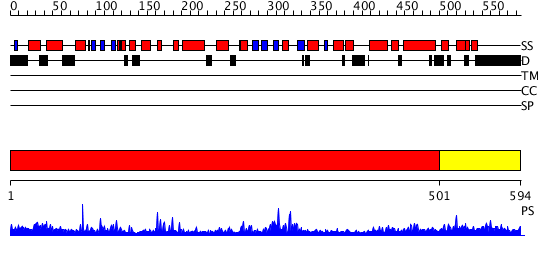
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..500] | 81.0 | RuBisCo LSMT C-terminal, substrate-binding domain; RuBisCo LSMT catalytic domain | |
| 2 | View Details | [501..594] | 1.04397 | View MSA. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.94 |
Source: Reynolds et al. (2008)