













| General Information: |
|
| Name(s) found: |
SODF3_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 13 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 263 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleoid
[IDA]
chloroplast [IDA] |
| Biological Process: |
removal of superoxide radicals
[IC]
|
| Molecular Function: |
superoxide dismutase activity
[IDA]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSSCVVTTSC FYTISDSSIR LKSPKLLNLS NQQRRRSLRS RGGLKVEAYY GLKTPPYPLD 60
61 ALEPYMSRRT LEVHWGKHHR GYVDNLNKQL GKDDRLYGYT MEELIKATYN NGNPLPEFNN 120
121 AAQVYNHDFF WESMQPGGGD TPQKGVLEQI DKDFGSFTNF REKFTNAALT QFGSGWVWLV 180
181 LKREERRLEV VKTSNAINPL VWDDIPIICV DVWEHSYYLD YKNDRAKYIN TFLNHLVSWN 240
241 AAMSRMARAE AFVNLGEPNI PIA |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
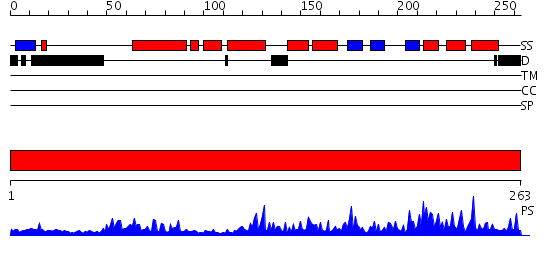
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..263] | 71.221849 | The crystal structure of the eukaryotic FeSOD from Vigna unguiculata suggests a new enzymatic mechanism |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.86 |
Source: Reynolds et al. (2008)