













| General Information: |
|
| Name(s) found: |
gi|9758896
[NCBI NR]
gi|28059343 [NCBI NR] gi|20260574 [NCBI NR] gi|15238746 [NCBI NR] |
| Description(s) found:
Found 13 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 333 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
protein folding
[IEA][ISS]
|
| Molecular Function: |
heat shock protein binding
[IEA]
unfolded protein binding [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MQTHLLVGPI PLKGYRRFSS SSFSGDLLPP SSNPIGRDLF PHRRRHRDGK SRSYRNRSKT 60
61 TITSAAFSSS SNTGGQNHYA VLGIARNATQ GDIKRAYRLL ARKFHPDVNK DSKAGELFKS 120
121 VRCSYEVLSN EATRTQYDRA LKLQENSRFH RVKRHSYTPE VEDAMKYYYT WSEKRQRSRH 180
181 GRFYGHYSTY PNSHFYAETE PQEEEGVETA PDQRDSFVEA LKSAFLSMFL LYTFGCLASL 240
241 TFSTFTALLD KELDMGYKVG FMIAWILGGK GGILLTLCLT FASWLCGKAS SSVVVLVLVA 300
301 MWVGSNLARH APLPQGALLT LLYMSIKLQV DST |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
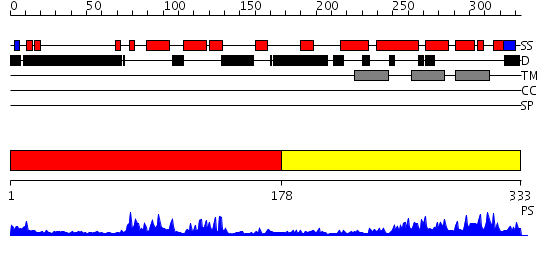
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..177] | 27.69897 | NMR STRUCTURE OF THE J-DOMAIN (RESIDUES 2-76) IN THE ESCHERICHIA COLI N-TERMINAL FRAGMENT (RESIDUES 2-108) OF THE MOLECULAR CHAPERONE DNAJ, 20 STRUCTURES | |
| 2 | View Details | [178..333] | 13.0 | The crystal structure of Hsp40 Ydj1 |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||
| 1 | No functions predicted. | ||||||
| 2 |
|
| Source: Reynolds et al. 2008. Manuscript submitted |

|