













| General Information: |
|
| Name(s) found: |
TPPF_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 14 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 368 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
metabolic process
[IEA]
trehalose biosynthetic process [IEA][ISS] |
| Molecular Function: |
catalytic activity
[IEA]
trehalose-phosphatase activity [ISS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MDLNSNHKSS VLKDPSPSVN QSRLGVSSRF MMSQWKKPAK LDDVRSNGWL DAMISSSPPR 60
61 KKLVKDFNVE VAPEDDFAQR AWMVKYPSAI SSFAHIAAQA KKKKIAVFLD YDGTLSPIVD 120
121 DPDRAIMSDA MRSAVKDVAS YFPTAIISGR SRDKVYQLVG LTELYYAGSH GMDIMTSSDG 180
181 PNCFKSTDQQ GKEVNLFQPA REFIPVIDEV FRTLVEKMKD IKGAKVENHK FCASVHYRNV 240
241 DEKDWPIIAQ RVHDHLKQYP RLRLTHGRKV LEVRPVIDWN KGRAVEFLLE SLGLSNKDDL 300
301 LPIYIGDDTT DEDAFKVLRD GNRGFGILVS SIPKESNAFY SLRDPSEVKK FLKTLVKWAK 360
361 LEKNSTGF |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
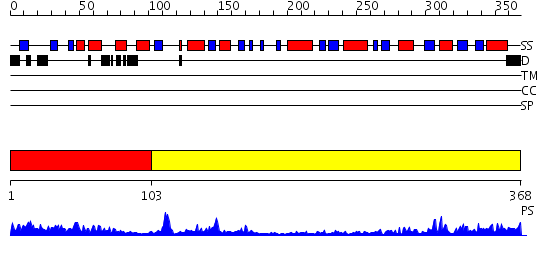
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..102] | 17.154902 | Trehalose-6-phosphate from E. coli bound with UDP-2-fluoro glucose. | |
| 2 | View Details | [103..368] | 33.221849 | Crystal structure of NYSGRC target T1436: A Hypothetical protein yidA. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||
| 1 | No functions predicted. | ||||||
| 2 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)