













| General Information: |
|
| Name(s) found: |
DRP3B_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 12 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 780 amino acids |
Gene Ontology: |
|
| Cellular Component: |
plasma membrane
[IDA]
|
| Biological Process: | NONE FOUND |
| Molecular Function: |
GTP binding
[IEA]
GTPase activity [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSVDDLPPSS ASAVTPLGSS VIPIVNKLQD IFAQLGSQST IALPQVAVVG SQSSGKSSVL 60
61 EALVGRDFLP RGNDICTRRP LRLQLVQTKP SSDGGSDEEW GEFLHHDPVR RIYDFSEIRR 120
121 EIEAETNRVS GENKGVSDIP IGLKIFSPNV LDISLVDLPG ITKVPVGDQP SDIEARIRTM 180
181 ILTYIKEPSC LILAVSPANT DLANSDALQI AGNADPDGHR TIGVITKLDI MDRGTDARNH 240
241 LLGKTIPLRL GYVGVVNRSQ EDILMNRSIK DALVAEEKFF RSRPVYSGLT DRLGVPQLAK 300
301 KLNQVLVQHI KALLPSLKSR INNALFATAK EYESYGDITE SRGGQGALLL SFITKYCEAY 360
361 SSTLEGKSKE MSTSELSGGA RILYIFQSVF VKSLEEVDPC EDLTADDIRT AIQNATGPRS 420
421 ALFVPDVPFE VLVRRQISRL LDPSLQCARF IFDELVKISH QCMMKELQRF PVLQKRMDEV 480
481 IGNFLREGLE PSQAMIRDLI EMEMDYINTS HPNFIGGTKA VEQAMQTVKS SRIPHPVARP 540
541 RDTVEPERTA SSGSQIKTRS FLGRQANGII TDQAVPTAAD AERPAPAGST SWSGFSSIFR 600
601 GSDGQAAAKN NLLNKPFSET TQEVYQNLST IYLKEPPTIL KSSETHSEQE SVEIEITKLL 660
661 LKSYYDIVRK NVEDLVPKAI MHFLVNYTKR ELHNVFIEKL YRENLIEELL KEPDELAIKR 720
721 KRTQETLRIL QQANRTLDEL PLEAESVERG YKIGSEAKHE ELPGTRRSRT ETNGNGRLHM 780
781 |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
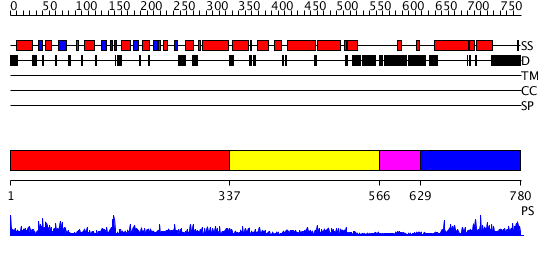
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..336] | 94.30103 | Structure of the nucleotide-free myosin II motor domain from Dictyostelium discoideum fused to the GTPase domain of dynamin 1 from Rattus norvegicus | |
| 2 | View Details | [337..565] | 2.372977 | View MSA. No confident structure predictions are available. | |
| 3 | View Details | [566..628] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [629..780] | 38.275724 | No description for PF02212.9 was found. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.96 |
Source: Reynolds et al. (2008)