













| General Information: |
|
| Name(s) found: |
PLCD7_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 12 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 584 amino acids |
Gene Ontology: |
|
| Cellular Component: |
plasma membrane
[IDA]
|
| Biological Process: |
intracellular signaling pathway
[IEA]
signal transduction [IEA][ISS] lipid metabolic process [IEA] |
| Molecular Function: |
phosphoinositide phospholipase C activity
[IEA]
phospholipase C activity [IEA][ISS] phosphoric diester hydrolase activity [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSKQTYKVCF CFRRRYRHTV SVAPAEIKTL FDNYSDKGLM TTDLLLRFLI DVQKQDKATK 60
61 EEAQDIVNAS SSLLHRNGLH LDAFFKYLFA VTNSPLSSLE VHQDMDAPLS HYFIYTGHNS 120
121 YLTGNQLSSD CSELPIIEAL KKGVRVIELD IWPNSDEDGI DVLHGRTLTS PVELIKCLRA 180
181 IREHAFDVSD YPVVVTLEDH LTPKLQAKVA EMVTDIFGEM LFTPPSGECL KEFPSPAFLK 240
241 KRIMISTKPP KEYKAATDDD LVKKGRDLGD KEVWGREVPS FIRRDRSVDK NDSNGDDDDD 300
301 DDDDDDDDDG DDKIKKNAPP EYKHLIAIEA GKPKGGITEC LKVDPDKVRR LSLSEEQLEK 360
361 ASEKYAKQIV RFTQRNLLRV YPKGTRITSS NYNPLIAWSH GAQMVAFNMQ GLGRSLWVMQ 420
421 GMFRGNGGCG YIKKPDLLLK SNAVFDPEAT LPVKTTLRVT IYMGEGWYYD FPHTHFDRYS 480
481 PPDFYTRVGI AGVPADTVMK KTKTLEDNWI PAWDEVFEFP LTVPELALLR IEVHEYDMSE 540
541 KDDFGGQICL PVWELRQGIR AVPLRNQDGV KCRSVKLLVR LEFV |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
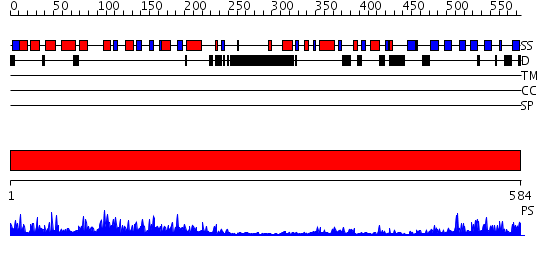
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..584] | 148.0 | Phosphoinositide-specific phospholipase C, isozyme D1 (PLC-D!); PI-specific phospholipase C isozyme D1 (PLC-D1), C-terminal domain; Phospholipase C isozyme D1 (PLC-D1) |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.95 |
Source: Reynolds et al. (2008)