













| General Information: |
|
| Name(s) found: |
SERA2_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 24 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 624 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IDA]
chloroplast [IDA] cytosol [IDA] |
| Biological Process: |
L-serine biosynthetic process
[TAS]
|
| Molecular Function: |
protein binding
[IPI]
phosphoglycerate dehydrogenase activity [IGI] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAFSSSCSSV KAVNSRWTSP SPSPSSRFAV LPAFLHRRYA TSVKLTAISA ALKTVEQTTL 60
61 TEDNRFSTVG SDSDEYNPTL PKPRILVTEK LGEAGVNLLR EFGDVDCSYD LSPEDLKKKV 120
121 AESDALIVRS GTKVTREVFE AAKGRLKVVG RAGVGIDNVD LQAATEHGCL VVNAPTANTV 180
181 AAAEHGIALL ASMARNVAQA DASIKAGKWE RSKYVGVSLV GKTLAVMGFG KVGTEVARRA 240
241 KGLGMTVISH DPYAPADRAR ALGVDLVSFD QAISTADFVS LHMPLTPATK KVFNDETFSK 300
301 MKKGVRLINV ARGGVIDEDA LVRALDAGIV AQAALDVFCE EPPSKDSRLI QHENVTVTPH 360
361 LGASTKEAQE GVAIEIAEAV AGALKGELSA TAVNAPMVAP EVLSELTPYI VLAEKLGRLA 420
421 VQLASGGKGV QSIRVVYRSA RDRDDLDTRL LRAMITKGII EPISDSYVNL VNADFIAKQK 480
481 GLRISEERMV VDSSPEYPVD SIQVQILNVE SNFAGAVSDA GDISIEGKVK YGVPHLTCVG 540
541 SFGVDVSLEG NLILCRQVDQ PGMIGQVGNI LGEQNVNVNF MSVGRTVLRK QAIMAIGVDE 600
601 EPDNKTLERI GGVSAIEEFV FLKL |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
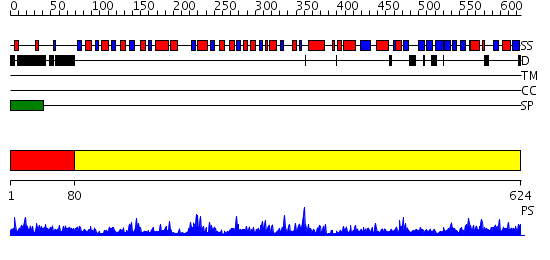
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..79] | 82.69897 | S-adenosylhomocystein hydrolase | |
| 2 | View Details | [80..624] | 110.0 | Crystal Structure of D-3-Phosphoglycerate dehydrogenase From Mycobacterium tuberculosis |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.75 |
Source: Reynolds et al. (2008)