













| General Information: |
|
| Name(s) found: |
PI3K_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 15 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 814 amino acids |
Gene Ontology: |
|
| Cellular Component: |
phosphoinositide 3-kinase complex
[IEA]
|
| Biological Process: |
oxygen and reactive oxygen species metabolic process
[IMP]
response to salt stress [IMP] microgametogenesis [IMP] endocytosis [IDA][IMP] |
| Molecular Function: |
inositol or phosphatidylinositol kinase activity
[IEA]
binding [IEA] phosphotransferase activity, alcohol group as acceptor [IEA] 1-phosphatidylinositol-3-kinase activity [IEA][ISS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MGANEFRFFL SCDINSPVTF RIEKLDGNLP VKKSSDSGVV SIAEEKKPEL YIECALYIDG 60
61 APFGLPMRTR LKTTGPPYCW NELITLSSKY RDLTAHSQLA ITVWDVSCGK TEGLIGGATV 120
121 LLFNSKMQMK SGKQKLRLWQ GKEADGSFPT STPGKVPRHE RGELERLEKL MNKYERGQIQ 180
181 SIDWLDRLML KSLDTIKEQE STKHGSSHLF VVIDFCSFEH RVVFQESGAN LFITAPIGST 240
241 NEFVTVWDTE LGKTNPSENK QLKLARSLDR GIIDRDLKPS NIERKSIQRV LKYPPTRTLS 300
301 GDERQLLWKF RFSLMSEKRA LTKFLRCVEW SDVQEAKQAI QLMYKWEMID VCDALELLSP 360
361 LFESEEVRAY AVSVLERADD EELQCYLLQL VQALRFERSD RSCLSQFLVQ RALQNIELAS 420
421 FLRWYVAVEL HDHVYAKRFY STYELLEENI IKLPPGVNGE DGYQLWQSLV RQTELTAQLC 480
481 SITREVRNVR GNTQKKIEKL RQLLGGLLSE LTYFEEPIRS PLTPNVLIKG IVAGESSLFK 540
541 SALHPLRLTF RTPEEGGSCK LIFKKGDDLR QDQLVVQMVW LMDRLLKLEN LDLCLTPYKV 600
601 LATGHDEGML EFIPSRSLAQ ILSEHRSITS YLQKFHPDEH APFGITATCL DTFIKSCAGY 660
661 SVITYILGIG DRHLDNLLLT DDGRLFHVDF AFILGRDPKP FPPPMKLCKE MVEAMGGAES 720
721 QYYTRFKSYC CEAYNILRKS SNLILNLFHL MAGSTIPDIA SDPEKGILKL QEKFRLDMDD 780
781 EACIHFFQDL INESVSALFP QMVETIHRWA QYWR |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
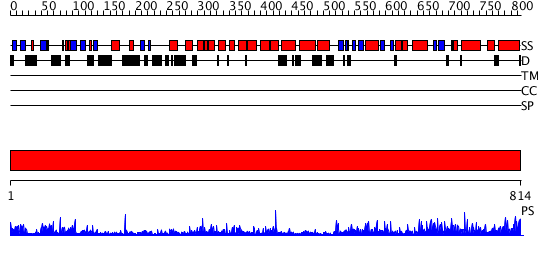
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..814] | 151.0 | No description for 2rd0A was found. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)