













| General Information: |
|
| Name(s) found: |
AROF_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 20 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 525 amino acids |
Gene Ontology: |
|
| Cellular Component: |
chloroplast
[TAS]
|
| Biological Process: |
aromatic amino acid family biosynthetic process
[TAS]
response to bacterium [IEP] response to wounding [IEP] chorismate biosynthetic process [IDA][IGI] |
| Molecular Function: |
protein binding
[IPI]
3-deoxy-7-phosphoheptulonate synthase activity [IDA][IGI][TAS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MALSNASSLS TRSIYGGDLS HRPSNRQSSF TFHPAVNTKP KSVNLVTAVH AAEPARNAVS 60
61 VKESVASSSS GALKWTPESW KLKKALQLPD YPNANELESV LKTIEAFPPI VFAGEARNLE 120
121 ERLADAAVGK AFLLQGGDCA ESFKEFNATN IRDTFRVLLQ MSIVLTFGGQ VPVIKVGRMA 180
181 GQFAKPRSDA FEEKDGVKLP SYKGDNINGD TFDEKSRIPD PNRMIRAYTQ SAATLNLLRA 240
241 FATGGYAAIQ RVTQWNLDFV EQSEQADRYQ ELANRVDEAL GFMSACGLGT DHPLMTTTDF 300
301 YTSHECLLLP YEQSLTRLDS TSGLYYDCSA HMVWCGERTR QLDGAHVEFL RGIANPLGIK 360
361 VSNKMDPFEL VKLVEILNPN NKPGRITVIV RMGAENMRVK LPHLIRAVRR SGQIVTWVCD 420
421 PMHGNTIKAP CGLKTRAFDS ILAEVRAFLD VHEQEGSHAG GIHLEMTGQN VTECIGGSRT 480
481 VTYDDLSSRY HTHCDPRLNA SQSLELAFIV AERLRKRRTG SQRVS |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
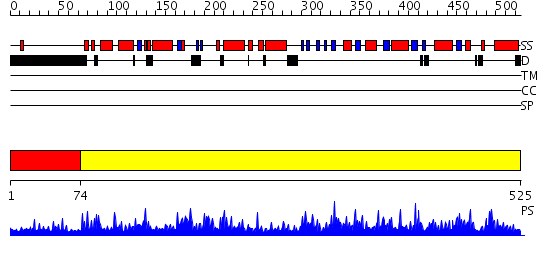
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..73] | 1.049994 | View MSA. No confident structure predictions are available. | |
| 2 | View Details | [74..525] | 1000.0 | The Structure of 3-Deoxy-D-Arabino-Heptulosonate 7-Phosphate Synthase from Mycobacterium tuberculosis |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||
| 1 | No functions predicted. | |||||||||||||||||||||||||||
| 2 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.92 |
Source: Reynolds et al. (2008)