













| General Information: |
|
| Name(s) found: |
CE34178
[WormBase]
|
| Description(s) found:
Found 15 descriptions. SHOW ALL |
|
| Organism: | Caenorhabditis elegans |
| Length: | 764 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
carbohydrate metabolic process
[IEA embryonic development ending in birth or egg hatching [IMP |
| Molecular Function: |
hydrolase activity, hydrolyzing O-glycosyl compounds
[IEA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MPEEYIMSSK ACIYVLGGAA NTAALFDFDG VYILDGGFAE KNPGFVAHVR DVSAVLLAAP 60
61 TLGNLGTTSA LLEQGKPLPV FTNTKPFKTA KPGSSGEIAK AIQEANSKIL SVAPPLFNPK 120
121 YPANIIYQSA AKGVLSLYIL AGDVKDAEVI TKALAGGNEA EVEKAAAEHG TIGVLLWRPA 180
181 MTDQSVVRVL ISGTSSLSRI QQSLDKAAKS LPFLNVPTVK SKDALSDIPA PAVPRPVAGK 240
241 PSARPATTTG TATRPTRPAV PAASAPRALT SRAPAGPSRP TTTRNAAPAP RTAVPSRATV 300
301 LTKTAPASKA PTRAPVPARS APAPPRGAPA KPAANTAKAE PIAQKKTVGK VQGTAPSKPA 360
361 PAAPASAATS PAPAPEAPRR DPNNVTIVLD DSLSPEDFNQ GSAPMDIVVI PPTPEPPRHE 420
421 VAQATHPEES IIDAEDLAKE PLIPQAPRDD GTLAECSEEV SKLVEISMDT DNSAEVAADL 480
481 AKAVGEVTQL SADLQNLGLD EKTDEYVRKL SNQMIEDATL PFTSALASSI VTSNGSETNG 540
541 HGEQAHAAQN GGIDHQKEIP KHDLMQSRSS VIENGAAVQY EKTDPALDDV LNACAQESEK 600
601 IDASHPDNLH MPAAPGSAAP AKPVKFARPY YFDVVTVPRN EKLETSVAAD GLQEFISKVR 660
661 SRNVILASKD ISGEQLQAIL CGKQTWCDSG KFENSFDLIN YFSAHPCNVI PTHSSPMLLD 720
721 FRQKNEEQFA ANNLQFSIPV EKQRTTVSSD AGAIEYELAR VDLL |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | hcp1 IP from december2004 | Cheeseman, I.M., et al. (2005) | |
| View Run | hcp2 ip from december | Cheeseman, I.M., et al. (2005) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
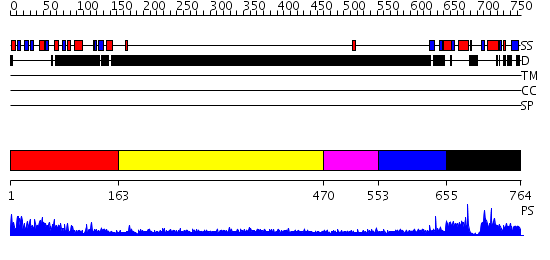
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..162] | 1.27 | Crystal structure of the complete modular teichioic acid phosphorylcholine esterase Pce (CbpE) from Streptococcus pneumoniae | |
| 2 | View Details | [163..469] | 3.257994 | View MSA. No confident structure predictions are available. | |
| 3 | View Details | [470..552] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [553..654] | 2.232991 | View MSA. No confident structure predictions are available. | |
| 5 | View Details | [655..764] | N/A | No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||
| 1 |
|
||||||
| 2 | No functions predicted. | ||||||
| 3 | No functions predicted. | ||||||
| 4 | No functions predicted. | ||||||
| 5 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.83 |
Source: Reynolds et al. (2008)