













| General Information: |
|
| Name(s) found: |
CE19598
[WormBase]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Caenorhabditis elegans |
| Length: | 798 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MLPSTSKSVE VNENSEEPIE LEENVIVVTG DEIQAEIAVV DGEEVIFADD DEYIEYDPLS 60
61 IANDRNAVYD IYQPHESENG TFMCKICMRT GKNTEYEDRS TFVAHRYKCH GSFNNNVMCP 120
121 LGDCREVFAS LYTLRRHLSQ QHELPIEIHL QSFVNIGEFE KFRHLVELAS GCRFMMHTKQ 180
181 PKYRRQVMHC SKSEHKLVLQ TQKHRLPRER MLKEGSACPS VISYRVESSS GEVHTFMQLY 240
241 HIGHAPDTES QNSNADNARP MDIVFPIKPA CWANHPMQYV QIDVHEMPTS IYGNKIYDNI 300
301 LIVTCLKTRF LWAKPLFECT RTAIGRILNS IFNEYGVPEG FSTSFSPTYI RDTIKSLESV 360
361 YAVEIREVWN EPLPYNCLER WVLELAQNEL GTRNRWVEQL QFLVMEYNQK PIPDRMETPF 420
421 ERMFNRKAPN LYHNGDNQEI IHDKYISYHS ELRNEEEGVE NRLNTSFEPG RKVFLRKGIT 480
481 KPRRGNNTQY YFGYIGEVDP SNPYYPFKVH YTSSDSPWPS ERNIYAWVSV FDLLPTIHEI 540
541 SEMSMTSKKV SIAGLLCSCM GIESSQKYDL TAMGDIARES SRCLLFRNCL CTNQMSRFCC 600
601 KLVGREHCRF HSVYPENDES YAKMDAALSS FIAYNKDSSK EAEKDLQDST PIEQILNEVG 660
661 RRRLESEITV EDPEDLAPPI LEPMEPTERH LDTEEEEEEP NQIPMASEHI DIEGVDEEEI 720
721 MEHSVNAISE EILFEYADDK RDDGPSTPKR GRRRKSPSES AGKKKKLSNN DEFVESTVSR 780
781 RSSARRTIRP KNLDDYVE |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | hcp2 ip from december | Cheeseman, I.M., et al. (2005) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
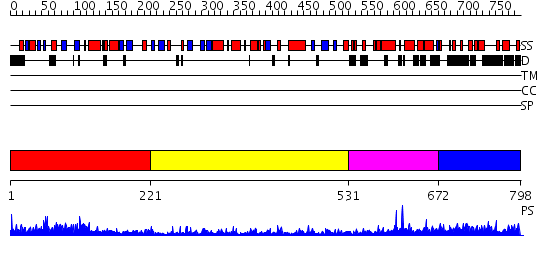
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..220] | 4.48299 | View MSA. No confident structure predictions are available. | |
| 2 | View Details | [221..530] | 4.2 | N-terminal Zn binding domain of HIV integrase; Retroviral integrase, catalytic domain | |
| 3 | View Details | [531..671] | 7.414995 | View MSA. No confident structure predictions are available. | |
| 4 | View Details | [672..798] | N/A | No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||
| 1 | No functions predicted. | |||||||||
| 2 |
|
|||||||||
| 3 | No functions predicted. | |||||||||
| 4 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)