













| General Information: |
|
| Name(s) found: |
ceh-26 /
CE00266
[WormBase]
|
| Description(s) found:
Found 18 descriptions. SHOW ALL |
|
| Organism: | Caenorhabditis elegans |
| Length: | 586 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IEA |
| Biological Process: |
molting cycle, protein-based cuticle
[IMP locomotory behavior [IMP multicellular organismal development [IEA nematode larval development [IMP locomotion [IMP positive regulation of growth rate [IMP regulation of transcription [IEA |
| Molecular Function: |
DNA binding
[IEA transcription regulator activity [IEA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSSGLPSIAA TAQPSANGFS FNNGFPPFGI YYSQLPPVNS NGSTPNSSNR FKVKRHRQRV 60
61 DAGEPRNTYQ GASTSVSSNS SSSSSTSNTN STPSSSSTSS KKSTEGMTET ETMTASIEQE 120
121 KVIQNEESEA GKDGMEEHDD GMNDFEIIDD TNDEVEESEE RESPESNSTL LQKGKAFNHR 180
181 KLERKTEQIN VTVGGTEEAE ETTDEAEGRE LSALQAQLQN GHIAEQQRRF YAAFLDQQRK 240
241 QVAAAQILHN GKINIERLVS SVKGDLVTNF MKDLEKVIRD WAADEVLKQE AAAPPQIPQP 300
301 PPQFHPIFPA NPLFHMAPPH HPVAGGGIFP PNFNAFNAFN ALRRNLQDVD TSSEMKKKRT 360
361 KVEIKKEDAM SSRASPLSAS ASPPLSRFFP APTMVGHYGG MNFGDREDSP TNSDELSECG 420
421 YEGGGSSSML TPMHLRKAKL MFFYTRYPNS NLLKSYFPDI RFNKNNTAQL VKWFSNFREF 480
481 YYNQMEKFAR QALAEGITDR NDIFVSKDSE LFKVLNTHYN RNNHIKAPDR LVFVVQETLR 540
541 EFHDAIKQGK DIEPSWKKTI YKVINRLEDQ IPDFFKEPNF LERLET |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
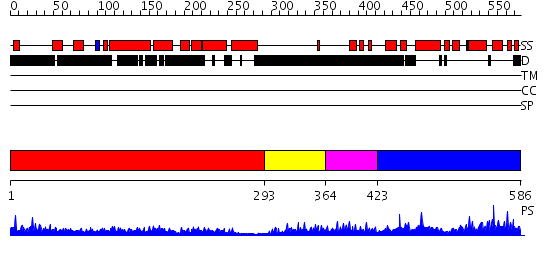
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..292] | 14.0 | Heavy meromyosin subfragment | |
| 2 | View Details | [293..363] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [364..422] | 2.115989 | View MSA. No confident structure predictions are available. | |
| 4 | View Details | [423..586] | 81.0 | Structural basis of prospero-DNA interaction; implications for transcription regulation in developing cells |
| Source: Reynolds et al. 2008. Manuscript submitted |

|