













| General Information: |
|
| Name(s) found: |
FBpp0306562
[FlyBase]
FBpp0302741 [FlyBase] Hrb87F-PC / FBpp0289817 [FlyBase] Hrb87F-PA / FBpp0082316 [FlyBase] |
| Description(s) found:
Found 45 descriptions. SHOW ALL |
|
| Organism: | Drosophila melanogaster |
| Length: | 385 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IC][IDA]
polytene chromosome puff [IDA] ribonucleoprotein complex [IDA][IPI][ISS][NAS] omega speckle [IDA] heterogeneous nuclear ribonucleoprotein complex [ISS] chromatin [IDA] nucleoplasm [IDA] |
| Biological Process: |
regulation of alternative nuclear mRNA splicing, via spliceosome
[IMP]
|
| Molecular Function: |
nucleotide binding
[IEA]
mRNA binding [ISS][NAS] sequence-specific DNA binding [IDA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAEQNDSNGN YDDGEEITEP EQLRKLFIGG LDYRTTDDGL KAHFEKWGNI VDVVVMKDPK 60
61 TKRSRGFGFI TYSQSYMIDN AQNARPHKID GRTVEPKRAV PRQEIDSPNA GATVKKLFVG 120
121 GLRDDHDEEC LREYFKDFGQ IVSVNIVSDK DTGKKRGFAF IEFDDYDPVD KIILQKTHSI 180
181 KNKTLDVKKA IAKQDMDRQG GGGGRGGPRA GGRGGQGDRG QGGGGWGGQN RQNGGGNWGG 240
241 AGGGGGFGNS GGNFGGGQGG GSGGWNQQGG SGGGPWNNQG GGNGGWNGGG GGGYGGGNSN 300
301 GSWGGNGGGG GGGGGFGNEY QQSYGGGPQR NSNFGNNRPA PYSQGGGGGG FNKGNQGGGQ 360
361 GFAGNNYNTG GGGQGGNMGG GNRRY |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
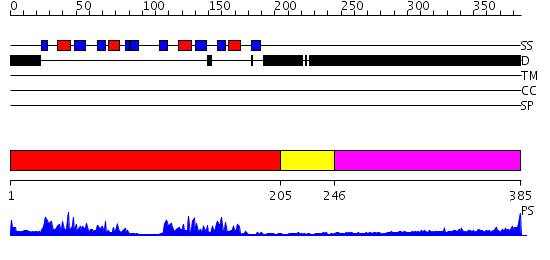
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..204] | 57.0 | Crystal Structure of UP1 Complexed With d(TTAGGGTTAG(6-MI)G); A Human Telomeric Repeat Containing 6-methyl-8-(2-deoxy-beta-ribofuranosyl)isoxanthopteridine (6-MI) | |
| 2 | View Details | [205..245] | 0.981 | Ribonuclease domain of colicin E3; Colicin E3 translocation domain; Colicin E3 receptor domain | |
| 3 | View Details | [246..385] | 3.09691 | Sec24 |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||
| 1 |
|
||||||
| 2 | No functions predicted. | ||||||
| 3 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)