













| General Information: |
|
| Name(s) found: |
ACTN4
[HGNC (HUGO)]
|
| Description(s) found:
Found 30 descriptions. SHOW ALL |
|
| Organism: | Homo sapiens |
| Length: | 911 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MVDYHAANQS YQYGPSSAGN GAGGGGSMGD YMAQEDDWDR DLLLDPAWEK QQRKTFTAWC 60
61 NSHLRKAGTQ IENIDEDFRD GLKLMLLLEV ISGERLPKPE RGKMRVHKIN NVNKALDFIA 120
121 SKGVKLVSIG AEEIVDGNAK MTLGMIWTII LRFAIQDISV EETSAKEGLL LWCQRKTAPY 180
181 KNVNVQNFHI SWKDGLAFNA LIHRHRPELI EYDKLRKDDP VTNLNNAFEV AEKYLDIPKM 240
241 LDAEDIVNTA RPDEKAIMTY VSSFYHAFSG AQKAETAANR ICKVLAVNQE NEHLMEDYEK 300
301 LASDLLEWIR RTIPWLEDRV PQKTIQEMQQ KLEDFRDYRR VHKPPKVQEK CQLEINFNTL 360
361 QTKLRLSNRP AFMPSEGKMV SDINNGWQHL EQAEKGYEEW LLNEIRRLER LDHLAEKFRQ 420
421 KASIHEAWTD GKEAMLKHRD YETATLSDIK ALIRKHEAFE SDLAAHQDRV EQIAAIAQEL 480
481 NELDYYDSHN VNTRCQKICD QWDALGSLTH SRREALEKTE KQLEAIDQLH LEYAKRAAPF 540
541 NNWMESAMED LQDMFIVHTI EEIEGLISAH DQFKSTLPDA DREREAILAI HKEAQRIAES 600
601 NHIKLSGSNP YTTVTPQIIN SKWEKVQQLV PKRDHALLEE QSKQQSNEHL RRQFASQANV 660
661 VGPWIQTKME EIGRISIEMN GTLEDQLSHL KQYERSIVDY KPNLDLLEQQ HQLIQEALIF 720
721 DNKHTNYTME HIRVGWEQLL TTIARTINEV ENQILTRDAK GISQEQMQEF RASFNHFDKD 780
781 HGGALGPEEF KACLISLGYD VENDRQGEAE FNRIMSLVDP NHSGLVTFQA FIDFMSRETT 840
841 DTDTADQVIA SFKVLAGDKN FITAEELRRE LPPDQAEYCI ARMAPYQGPD AVPGALDYKS 900
901 FSTALYGESD L |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | Sample pIC145 from april 2005 | Okada, M., et al. (2006) | |
| View Run | No Comments | Cheeseman, I.M., et al. (2004) |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
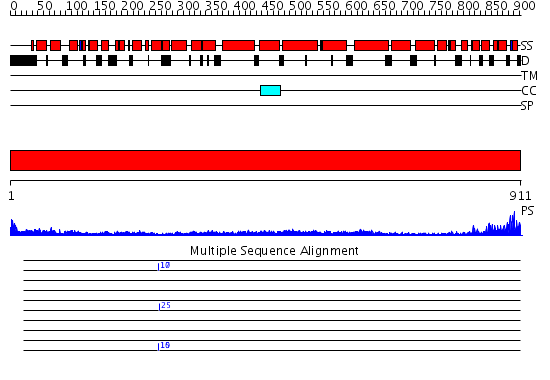
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..911] | 74.69897 | Cryo-EM Structure of Chicken Gizzard Smooth Muscle alpha-Actinin |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)