













| General Information: |
|
| Name(s) found: |
TCF3
[HGNC (HUGO)]
|
| Description(s) found:
Found 35 descriptions. SHOW ALL |
|
| Organism: | Homo sapiens |
| Length: | 654 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MNQPQRMAPV GTDKELSDLL DFSMMFPLPV TNGKGRPASL AGAQFGGSGL EDRPSSGSWG 60
61 SGDQSSSSFD PSRTFSEGTH FTESHSSLSS STFLGPGLGG KSGERGAYAS FGRDAGVGGL 120
121 TQAGFLSGEL ALNSPGPLSP SGMKGTSQYY PSYSGSSRRR AADGSLDTQP KKVRKVPPGL 180
181 PSSVYPPSSG EDYGRDATAY PSAKTPSSTY PAPFYVADGS LHPSAELWSP PGQAGFGPML 240
241 GGGSSPLPLP PGSGPVGSSG SSSTFGGLHQ HERMGYQLHG AEVNGGLPSA SSFSSAPGAT 300
301 YGGVSSHTPP VSGADSLLGS RGTTAGSSGD ALGKALASIY SPDHSSNNFS SSPSTPVGSP 360
361 QGLAGTSQWP RAGAPGALSP SYDGGLHGLQ SKIEDHLDEA IHVLRSHAVG TAGDMHTLLP 420
421 GHGALASGFT GPMSLGGRHA GLVGGSHPED GLAGSTSLMH NHAALPSQPG TLPDLSRPPD 480
481 SYSGLGRAGA TAAASEIKRE EKEDEENTSA ADHSEEEKKE LKAPRARTSP DEDEDDLLPP 540
541 EQKAEREKER RVANNARERL RVRDINEAFK ELGRMCQLHL NSEKPQTKLL ILHQAVSVIL 600
601 NLEQQVRERN LNPKAACLKR REEEKVSGVV GDPQMVLSAP HPGLSEAHNP AGHM |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | 3912: TAP-tagged, strain background includes hpc2(delta) | Green EM, et al (2005) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
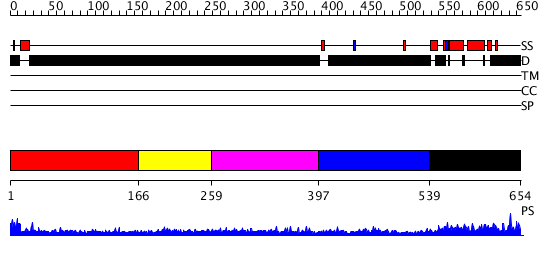
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..165] | 2.214992 | View MSA. No confident structure predictions are available. | |
| 2 | View Details | [166..258] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [259..396] | 5.238983 | View MSA. No confident structure predictions are available. | |
| 4 | View Details | [397..538] | N/A | No confident structure predictions are available. | |
| 5 | View Details | [539..654] | 2.02 | Myod B/HLH domain |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.97 |
Source: Reynolds et al. (2008)