













| General Information: |
|
| Name(s) found: |
VHS3_YEAST
[Swiss-Prot]
|
| Description(s) found: |
|
| Organism: | Saccharomyces cerevisiae S288c |
| Length: | 674 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MTNKSSLKNN RKGVASNTLS GAEQANIGSS AMPDTNSTGP FSSVSSLDTP VVRKSTSPTG 60
61 SQTKSIMNAS GTSGAVVSNT PEPGLKRIPT VTFSDPKLGS LRSDVEQTPP NQVARQSSEK 120
121 KATSVHIAAE GANQGRNLKD INTKVPKDGE ASASSFSTPT SILSNADMGN NISSLLAKKL 180
181 SFTGGTDSIL NSDNSSDSPR KEHPHFYVED PLHTPSVRSR SNSTSPRPSV VVNTFNPINI 240
241 EREGSISKTG EPTLLESVLE EAMSPNAVSN PLKRENIMTN MDPRLPQDDG KLHVLFGATG 300
301 SLSVFKLKHM IRKLEEIYGR DKICIQVILT NSATKFFAMK YMRKNKKQHN SIDTSFNSTN 360
361 SNAGNITGNK KKVASLEKFS IQKTSSNSAA SQTNNKQEEE KQMASTTGFP STLGGSRTYS 420
421 NSSNVVSQHP QIELPAHIQF WTDQDEWDVW RQRTDPVLHI ELRRWADILV VAPLTANTLA 480
481 KIALGLCDNL LTSVIRAWNP TFPIFLAPSM GSGTFNSIMT KKHFRIIQEE MPWVTVFKPS 540
541 EKVMGINGDI GLSGMMDANE IVGKIVVKLG GYPDVSAGKE EEEDEDNDEE DDNKKNDTGG 600
601 KDEDNDDDDD DDDDDDDDDD DDDDDDDDDD DDDDDDDDDD DDDDDDDDDD EDDEDEDEDD 660
661 EGKKKEDKGG LQRS |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
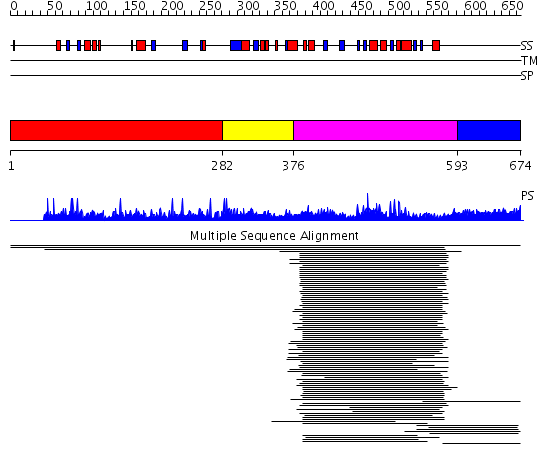
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..281] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [282..375] | 1.88 | 4'-phosphopantothenoylcysteine decarboxylase (PPC decarboxylase, halotolerance protein Hal3a) | |
| 3 | View Details | [376..592] | 371.9897 | 4'-phosphopantothenoylcysteine decarboxylase (PPC decarboxylase, halotolerance protein Hal3a) | |
| 4 | View Details | [593..674] | N/A | No confident structure predictions are available. | |
| 5 | View Details | [574..674] | 4.69897 | Strucuture of the C-terminal domain of human thrombospondin-2 |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||
| 1 | No functions predicted. | ||||||||||||||||||||||||
| 2 | No functions predicted. | ||||||||||||||||||||||||
| 3 | No functions predicted. | ||||||||||||||||||||||||
| 4 |
|
||||||||||||||||||||||||
| 5 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.71 |
Source: Reynolds et al. (2008)