













| General Information: |
|
| Name(s) found: |
CYK3_YEAST
[Swiss-Prot]
|
| Description(s) found: |
|
| Organism: | Saccharomyces cerevisiae S288c |
| Length: | 885 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MATNLTSLKP PFKVKARYGW SGQTKGDLGF LEGDIMEVTR IAGSWFYGKL LRNKKCSGYF 60
61 PHNFVILLEE RLNSSTENGR QPSKIVESFE KSNKVVIPPV PSRYSDERPR PKKKLSSSMP 120
121 NSPKKPVDSL TKARKAKSKE MVNEKNIYNT QSSRHHNNSA PNLPLASHSK PQVRNFEESM 180
181 NNPLPPLPPL PDLDNMRKTD KRAPKKSYSA NDLHMARSSR EYNYYKDNQK FYDGFIPEKR 240
241 YSLEEDSISS GLFSNSQYLN DSACSSENSF ALMSDFSATS AGSFARHKYA QSFSDSLQRS 300
301 QNANGCSTKI NDSQEFGDSN ASSRNGKMGD ILRKIIIPKR NTNIYSSSVS SPKSPKAYPK 360
361 LPDIQNLNLS ATPDEARDWI AVKCHLNRAR TLTKYDKHPR YMRALEENRD LILHPQDSIY 420
421 NGLNTNEVKG NTKPGLVDVE LAELNIEYID KMTWKRCIRD GTMTLDSWAQ TTFSARYSTV 480
481 LEKLRGIYIF CTEMFALTDD NGTSDFSAEP QNLEKILYRK HCTPYELTWL FKKLANSLGI 540
541 TCEIVIGFLK TPSAINWEFK YNHCWLRILV NKEWRFIDVI LGNVTNPIHE FVNNRKIKKA 600
601 ENSYFLMAPL EMIYTHIPPR EFEQHIVPSI DQLSALYLPL VFPSFFKNEL KLYKFSTALS 660
661 FLEDSEIYEC SLEIPNDVEV FASVVIPTDN EEASSAYRNM ELALTQIKKQ KAESGRRIAL 720
721 IKAVLPPNVN KGSLYIHSGV RGTQTSIANI HPLSMMVPLT HKGSNMKYEF VIKIPSESIQ 780
781 KIELYIVEPQ SRYLFVGNEY SFEVIQSPSD GIVYSSDEGP NQNRKQPMAI KSPSGRVHEL 840
841 VKSDPHFPYG TWKGSIKIKE PGVWSALVIA DSGIGWSVFA EWLCV |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
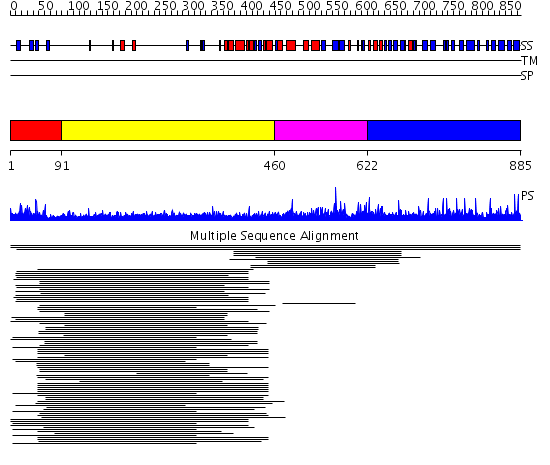
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..90] | 5.69897 | Carboxyl-terminal src kinase (csk) | |
| 2 | View Details | [91..459] | 28.488997 | View MSA. No confident structure predictions are available. | |
| 3 | View Details | [460..621] | 8.13 | Arylamine N-acetyltransferase | |
| 4 | View Details | [622..885] | N/A | No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||
| 1 | No functions predicted. | ||||||||||||
| 2 |
|
||||||||||||
| 3 | No functions predicted. | ||||||||||||
| 4 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.92 |
Source: Reynolds et al. (2008)