













| General Information: |
|
| Name(s) found: |
KN4A_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 13 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 1035 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytoplasm
[IDA]
|
| Biological Process: |
cellulose microfibril organization
[IMP]
|
| Molecular Function: |
microtubule motor activity
[ISS]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MESTPPPDDC SVKVAVHIRP LIGDERIQGC QDCVTVVTGK PQVQIGSHSF TFDHVYGSSG 60
61 SPSTEMYEEC AAPLVDGLFQ GYNATVLAYG QTGSGKTYTM GTGCGDSSQT GIIPQVMNAL 120
121 FTKIETLKQQ IEFQIHVSFI EIHKEEVQDL LDPCTVNKSD TNNTGHVGKV AHVPGKPPIQ 180
181 IRETSNGVIT LAGSTEVSVS TLKEMAACLD QGSVSRATGS TNMNNQSSRS HAIFTITVEQ 240
241 MRKINTDSPE NGAYNGSLKE EYLCAKLHLV DLAGSERAKR TGSDGLRFKE GVHINKGLLA 300
301 LGNVISALGD EKKRKDGAHV PYRDSKLTRL LQDSLGGNSR TVMIACISPA DINAEETLNT 360
361 LKYANRARNI RNKPVVNRDP VSSEMLKMRQ QVEYLQAELS LRTGGSSCAE VQALKERIVW 420
421 LETANEELCR ELHEYRSRCP GVEHSEKDFK DIRADDIVGS VRPDGLKRSL HSIESSNYPM 480
481 VEATTGDSRE IDEEAKEWEH KLLQNSMDKE LYELNRRLEE KESEMKLFDG YDPAALKQHF 540
541 GKKIAEVEDE KRSVQEERNR LLAEIENLAS DGQAQKLQDV HAQNLKALEA QILDLKKKQE 600
601 SQVQLLKQKQ KSDDAARRLQ DEIQSIKAQK VQLQHRMKQE AEQFRQWKAS REKELLQLRK 660
661 EGRKSEYERH KLQALNQRQK MVLQRKTEEA AMATKRLKEL LEARKSSPRE HSAGTNGFGT 720
721 NGQTNEKSLQ RWLDHELEVM VNVHEVRHEY EKQSHVRAAL AEELAVLRQV DEFAVKGLSP 780
781 PRGKNGFARA SSLSPNARMA RISSLENMLV ISSNSLVAMA SQLSEAEERE RAFTNRGRWN 840
841 QLRSMGEAKN LLQYMFNSLA ETRCQLWEKD VEIKEMKDQF KEIVGLLRQS ELRRKEAEKE 900
901 LKLREQAIAT SLGTPPSSVK HVAEDLSTPS PMTVPAQKQL KFTPGIANGK VRGPAAFLDT 960
961 NKKMVPMGQV SMRKLSAVGK QGGRLWRWKR SHHQWIVQFK WKWQKPWRLS EWIRTSDETL1020
1021 LKSKPRLKAL PNKIM |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
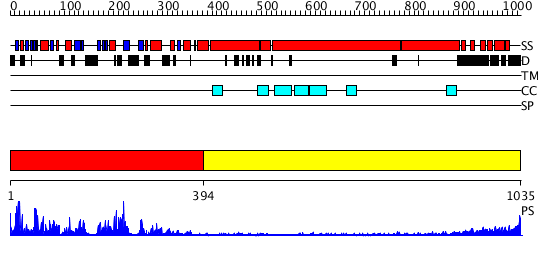
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..393] | 96.0 | No description for 3b6uA was found. | |
| 2 | View Details | [394..1035] | 9.522879 | No description for 2dfsA was found. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)