













| General Information: |
|
| Name(s) found: |
ARC5_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 7 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 777 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAEVSAKSVT VEEMAEEDDA AIEERWSLYE AYNELHALAQ ELETPFEAPA VLVVGQQTDG 60
61 KSALVEALMG FQFNHVGGGT KTRRPITLHM KYDPQCQFPL CHLGSDDDPS VSLPKSLSQI 120
121 QAYIEAENMR LEQEPCSPFS AKEIIVKVQY KYCPNLTIID TPGLIAPAPG LKNRALQVQA 180
181 RAVEALVRAK MQHKEFIILC LEDSSDWSIA TTRRIVMQVD PELSRTIVVS TKLDTKIPQF 240
241 SCSSDVEVFL SPPASALDSS LLGDSPFFTS VPSGRVGYGQ DSVYKSNDEF KQAVSLREME 300
301 DIASLEKKLG RLLTKQEKSR IGISKLRLFL EELLWKRYKE SVPLIIPLLG KEYRSTVRKL 360
361 DTVSKELSSL DEAKLKERGR TFHDLFLTKL SLLLKGTVVA PPDKFGETLQ DERTQGGAFV 420
421 GTDGLQFSHK LIPNAGMRLY GGAQYHRAMA EFRFLVGAIK CPPITREEIV NACGVEDIHD 480
481 GTNYSRTACV IAVAKARETF EPFLHQLGAR LLHILKRLLP ISVYLLQKEG EYLSGHEVFL 540
541 KRVASAFNSF VESTEKSCRD KCMEDLASTT RYVTWSLHNK NRAGLRQFLD SFGGTEHNTT 600
601 SGNAIGFSLP QDALGGTTDT KSRSDVKLSH LASNIDSGSS IQTTEMRLAD LLDSTLWNRK 660
661 LAPSSERIVY ALVQQIFQGI REYFLASAEL KFNCFLLMPI VDKLPALLRE ELENAFEDDL 720
721 DSIFDITNLR QSLDQKKRST EIELRRIKRI KEKFRVMNEK LNSHEFAQNL KAPSVQH |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
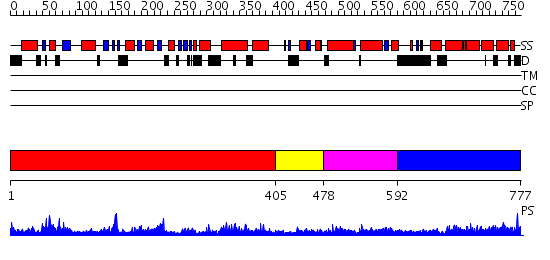
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..404] | 83.69897 | Dynamin G domain | |
| 2 | View Details | [405..477] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [478..591] | 2.118994 | View MSA. No confident structure predictions are available. | |
| 4 | View Details | [592..777] | 3.095996 | View MSA. No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||
| 1 |
|
|||||||||
| 2 | No functions predicted. | |||||||||
| 3 | No functions predicted. | |||||||||
| 4 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)