













| General Information: |
|
| Name(s) found: |
ZRAR_ECOLI
[Swiss-Prot]
|
| Description(s) found:
Found 7 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli |
| Length: | 441 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MTHDNIDILV VDDDISHCTI LQALLRGWGY NVALANSGRQ ALEQVREQVF DLVLCDVRMA 60
61 EMDGIATLKE IKALNPAIPV LIMTAYSSVE TAVEALKTGA LDYLIKPLDF DNLQATLEKA 120
121 LAHTHSIDAE TPAVTASQFG MVGKSPAMQH LLSEIALVAP SEATVLIHGD SGTGKELVAR 180
181 AIHASSARSE KPLVTLNCAA LNESLLESEL FGHEKGAFTG ADKRREGRFV EADGGTLFLD 240
241 EIGDISPMMQ VRLLRAIQER EVQRVGSNQI ISVDVRLIAA THRDLAAEVN AGRFRQDLYY 300
301 RLNVVAIEVP SLRQRREDIP LLAGHFLQRF AERNRKAVKG FTPQAMDLLI HYDWPGNIRE 360
361 LENAVERAVV LLTGEYISER ELPLAIASTP IPLGQSQDIQ PLVEVEKEVI LAALEKTGGN 420
421 KTEAARQLGI TRKTLLAKLS R |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
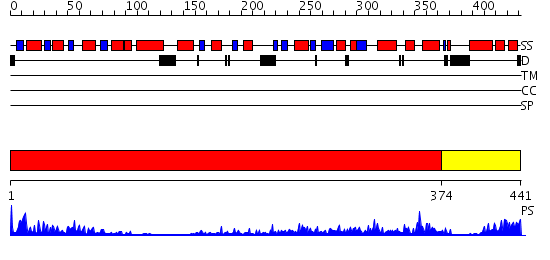
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..373] | 97.09691 | Crystal structure of sigm54 activator (AAA+ ATPase) in the inactive state | |
| 2 | View Details | [374..441] | 94.0 | Crystal structure of a sigma54-activator suggests the mechanism for the conformational switch necessary for sigma54 binding |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)