













| General Information: |
|
| Name(s) found: |
OXC_ECOLI
[Swiss-Prot]
|
| Description(s) found:
Found 20 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli |
| Length: | 564 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSDQLQMTDG MHIIVEALKQ NNIDTIYGVV GIPVTDMARH AQAEGIRYIG FRHEQSAGYA 60
61 AAASGFLTQK PGICLTVSAP GFLNGLTALA NATVNGFPMI MISGSSDRAI VDLQQGDYEE 120
121 LDQMNAAKPY AKAAFRVNQP QDLGIALARA IRVSVSGRPG GVYLDLPANV LAATMEKDEA 180
181 LTTIVKVENP SPALLPCPKS VTSAISLLAK AERPLIILGK GAAYSQADEQ LREFIESAQI 240
241 PFLPMSMAKG ILEDTHPLSA AAARSFALAN ADVVMLVGAR LNWLLAHGKK GWAADTQFIQ 300
301 LDIEPQEIDS NRPIAVPVVG DIASSMQGML AELKQNTFTT PLVWRDILNI HKQQNAQKMH 360
361 EKLSTDTQPL NYFNALSAVR DVLRENQDIY LVNEGANTLD NARNIIDMYK PRRRLDCGTW 420
421 GVMGIGMGYA IGASVTSGSP VVAIEGDSAF GFSGMEIETI CRYNLPVTIV IFNNGGIYRG 480
481 DGVDLSGAGA PSPTDLLHHA RYDKLMDAFR GVGYNVTTTD ELRHALTTGI QSRKPTIINV 540
541 VIDPAAGTES GHITKLNPKQ VAGN |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
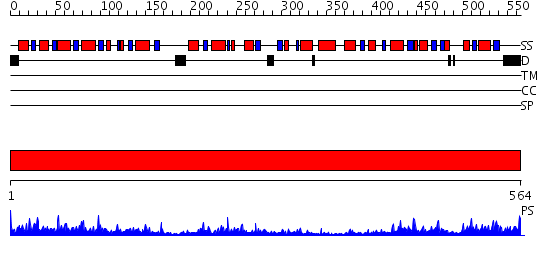
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..564] | 107.0 | CRYSTAL STRUCTURE OF OXALYL-COA DECARBOXYLASE IN COMPLEX WITH THE COFACTOR DERIVATIVE THIAMIN-2-THIAZOLONE DIPHOSPHATE AND ADENOSINE DIPHOSPHATE |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.63 |
Source: Reynolds et al. (2008)