













| General Information: |
|
| Name(s) found: |
MALT_ECOLI
[Swiss-Prot]
|
| Description(s) found:
Found 7 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli |
| Length: | 901 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MLIPSKLSRP VRLDHTVVRE RLLAKLSGAN NFRLALITSP AGYGKTTLIS QWAAGKNDIG 60
61 WYSLDEGDNQ QERFASYLIA AVQQATNGHC AICETMAQKR QYASLTSLFA QLFIELAEWH 120
121 SPLYLVIDDY HLITNPVIHE SMRFFIRHQP ENLTLVVLSR NLPQLGIANL RVRDQLLEIG 180
181 SQQLAFTHQE AKQFFDCRLS SPIEAAESSR ICDDVSGWAT ALQLIALSAR QNTHSAHKSA 240
241 RRLAGINASH LSDYLVDEVL DNVDLATRHF LLKSAILRSM NDALITRVTG EENGQMRLEE 300
301 IERQGLFLQR MDDTGEWFCY HPLFGNFLRQ RCQWELAAEL PEIHRAAAES WMAQGFPSEA 360
361 IHHALAAGDA LMLRDILLNH AWSLFNHSEL SLLEESLKAL PWDSLLENPQ LVLLQAWLMQ 420
421 SQHRYGEVNT LLARAEHEIK DIREDTMHAE FNALRAQVAI NDGNPDEAER LAKLALEELP 480
481 PGWFYSRIVA TSVLGEVLHC KGELTRSLAL MQQTEQMARQ HDVWHYALWS LIQQSEILFA 540
541 QGFLQTAWET QEKAFQLINE QHLEQLPMHE FLVRIRAQLL WAWARLDEAE ASARSGIEVL 600
601 SSYQPQQQLQ CLAMLIQCSL ARGDLDNARS QLNRLENLLG NGKYHSDWIS NANKVRVIYW 660
661 QMTGDKAAAA NWLRHTAKPE FANNHFLQGQ WRNIARAQIL LGEFEPAEIV LEELNENARS 720
721 LRLMSDLNRN LLLLNQLYWQ AGRKSDAQRV LLDALKLANR TGFISHFVIE GEAMAQQLRQ 780
781 LIQLNTLPEL EQHRAQRILR EINQHHRHKF AHFDENFVER LLNHPEVPEL IRTSPLTQRE 840
841 WQVLGLIYSG YSNEQIAGEL EVAATTIKTH IRNLYQKLGV AHRQDAVQHA QQLLKMMGYG 900
901 V |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
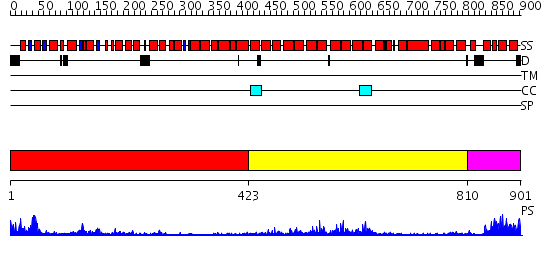
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..422] | 13.69897 | Structure of the apoptotic protease-activating factor 1 bound to ADP | |
| 2 | View Details | [423..809] | 16.69897 | Transcription factor MalT domain III | |
| 3 | View Details | [810..901] | 15.39794 | Crystallographic structure of response regulator StyR from Pseudomonas fluorescens |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||
| 1 | No functions predicted. | |||||||||||||||||||||||||||
| 2 |
|
|||||||||||||||||||||||||||
| 3 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.96 |
Source: Reynolds et al. (2008)