













| General Information: |
|
| Name(s) found: |
GYRA_ECOLI
[Swiss-Prot]
|
| Description(s) found:
Found 10 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli |
| Length: | 875 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSDLAREITP VNIEEELKSS YLDYAMSVIV GRALPDVRDG LKPVHRRVLY AMNVLGNDWN 60
61 KAYKKSARVV GDVIGKYHPH GDSAVYDTIV RMAQPFSLRY MLVDGQGNFG SIDGDSAAAM 120
121 RYTEIRLAKI AHELMADLEK ETVDFVDNYD GTEKIPDVMP TKIPNLLVNG SSGIAVGMAT 180
181 NIPPHNLTEV INGCLAYIDD EDISIEGLME HIPGPDFPTA AIINGRRGIE EAYRTGRGKV 240
241 YIRARAEVEV DAKTGRETII VHEIPYQVNK ARLIEKIAEL VKEKRVEGIS ALRDESDKDG 300
301 MRIVIEVKRD AVGEVVLNNL YSQTQLQVSF GINMVALHHG QPKIMNLKDI IAAFVRHRRE 360
361 VVTRRTIFEL RKARDRAHIL EALAVALANI DPIIELIRHA PTPAEAKTAL VANPWQLGNV 420
421 AAMLERAGDD AARPEWLEPE FGVRDGLYYL TEQQAQAILD LRLQKLTGLE HEKLLDEYKE 480
481 LLDQIAELLR ILGSADRLME VIREELELVR EQFGDKRRTE ITANSADINL EDLITQEDVV 540
541 VTLSHQGYVK YQPLSEYEAQ RRGGKGKSAA RIKEEDFIDR LLVANTHDHI LCFSSRGRVY 600
601 SMKVYQLPEA TRGARGRPIV NLLPLEQDER ITAILPVTEF EEGVKVFMAT ANGTVKKTVL 660
661 TEFNRLRTAG KVAIKLVDGD ELIGVDLTSG EDEVMLFSAE GKVVRFKESS VRAMGCNTTG 720
721 VRGIRLGEGD KVVSLIVPRG DGAILTATQN GYGKRTAVAE YPTKSRATKG VISIKVTERN 780
781 GLVVGAVQVD DCDQIMMITD AGTLVRTRVS EISIVGRNTQ GVILIRTAED ENVVGLQRVA 840
841 EPVDEEDLDT IDGSAAEGDD EIAPEVDVDD EPEEE |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
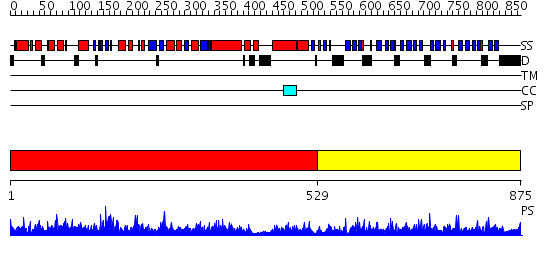
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..528] | 1000.0 | 59KDA FRAGMENT OF GYRASE A FROM E. COLI | |
| 2 | View Details | [529..875] | 108.0 | A Superhelical Spiral in Escherichia coli DNA Gyrase A C-terminal Domain Imparts Unidirectional Supercoiling Bias |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)