













| General Information: |
|
| Name(s) found: |
GLGB_ECOLI
[Swiss-Prot]
|
| Description(s) found:
Found 9 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli |
| Length: | 728 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSDRIDRDVI NALIAGHFAD PFSVLGMHKT TAGLEVRALL PDATDVWVIE PKTGRKLAKL 60
61 ECLDSRGFFS GVIPRRKNFF RYQLAVVWHG QQNLIDDPYR FGPLIQEMDA WLLSEGTHLR 120
121 PYETLGAHAD TMDGVTGTRF SVWAPNARRV SVVGQFNYWD GRRHPMRLRK ESGIWELFIP 180
181 GAHNGQLYKY EMIDANGNLR LKSDPYAFEA QMRPETASLI CGLPEKVVQT EERKKANQFD 240
241 APISIYEVHL GSWRRHTDNN FWLSYRELAD QLVPYAKWMG FTHLELLPIN EHPFDGSWGY 300
301 QPTGLYAPTR RFGTRDDFRY FIDAAHAAGL NVILDWVPGH FPTDDFALAE FDGTNLYEHS 360
361 DPREGYHQDW NTLIYNYGRR EVSNFLVGNA LYWIERFGID ALRVDAVASM IYRDYSRKEG 420
421 EWIPNEFGGR ENLEAIEFLR NTNRILGEQV SGAVTMAEES TDFPGVSRPQ DMGGLGFWYK 480
481 WNLGWMHDTL DYMKLDPVYR QYHHDKLTFG ILYNYTENFV LPLSHDEVVH GKKSILDRMP 540
541 GDAWQKFANL RAYYGWMWAF PGKKLLFMGN EFAQGREWNH DASLDWHLLE GGDNWHHGVQ 600
601 RLVRDLNLTY RHHKAMHELD FDPYGFEWLV VDDKERSVLI FVRRDKEGNE IIVASNFTPV 660
661 PRHDYRFGIN QPGKWREILN TDSMHYHGSN AGNGGTVHSD EIASHGRQHS LSLTLPPLAT 720
721 IWLVREAE |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
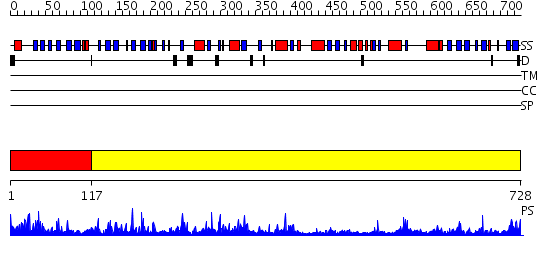
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..116] | 2.39 | 1,4-alpha-glucan branching enzyme, N-terminal domain N; 1,4-alpha-glucan branching enzyme; 1,4-alpha-glucan branching enzyme, central domain | |
| 2 | View Details | [117..728] | 157.0 | 1,4-alpha-glucan branching enzyme, N-terminal domain N; 1,4-alpha-glucan branching enzyme; 1,4-alpha-glucan branching enzyme, central domain |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 2 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.96 |
Source: Reynolds et al. (2008)