













| General Information: |
|
| Name(s) found: |
cct-7 /
CE28255
[WormBase]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Caenorhabditis elegans |
| Length: | 535 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
locomotory behavior
[IMP hermaphrodite genitalia development [IMP nematode larval development [IMP embryonic development ending in birth or egg hatching [IMP cellular protein metabolic process [IEA protein folding [IEA reproduction [IMP growth [IMP |
| Molecular Function: |
ATP binding
[IEA protein binding [IEA unfolded protein binding [IEA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MMRPPIILLK DGTENKQGKG QIISNINACQ VVADSIRTTL GPRGLDKLIV DSKGATTISN 60
61 DGATILKLLD IVFPAASTMV DIARSQDAEV GDGTTSVVVL AAEILKQMKP FIEDGVHPQL 120
121 LIRAIGKACE KTLKNLADLE IKINGETELR EMLVKCAATT LSSKLVSQER LFFANMIVDA 180
181 VNTLDAHLPL NMIGIKKVNG GNLHESRLIK GVAFQKAFSY AGFEMQPKKY SNVKVALLNI 240
241 ELELKAEKEN AEMRLTNVSD FQAVVDAEWN ILYDKLQKIH DSGANVVLSK LPIGDVATQW 300
301 FADRDMFCAG RIPQDDLDRL MSACGGSVLT TVSQIDDSVL GKCGKFYEQQ VGSERYNFFE 360
361 DCSKAQACTL LLRGGAEQFI AETERSLHDA IMIVRRAKKN DSIVAGGGAI EMELSRLIRE 420
421 HSKGIEGKDQ AFWMAYGQAF EIIPRQLCQN AGLDALDVLN KLRHRHAQGE KWAGIDIHRE 480
481 AVGDNLVACV WEPSIVKRNA ITAATEAACL VLSIDQTVKN VRTPSGGAMP GLPGQ |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | hcp1 IP from december2004 | Cheeseman, I.M., et al. (2005) |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
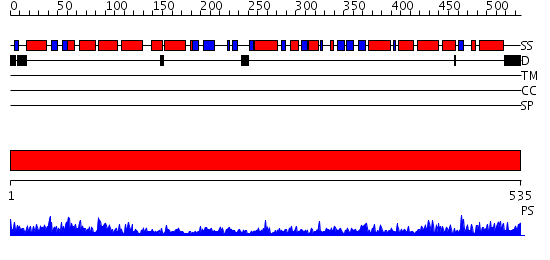
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..535] | 135.0 | Crystal structure of the chaperonin from Thermococcus strain KS-1 (nucleotide-free form of single mutant) |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)