













| General Information: |
|
| Name(s) found: |
CE27760
[WormBase]
|
| Description(s) found:
Found 23 descriptions. SHOW ALL |
|
| Organism: | Caenorhabditis elegans |
| Length: | 600 amino acids |
Gene Ontology: |
|
| Cellular Component: |
molybdopterin synthase complex
[IEA |
| Biological Process: |
locomotory behavior
[IMP locomotion [IMP positive regulation of growth rate [IMP determination of adult lifespan [IMP pyrroloquinoline quinone biosynthetic process [IEA Mo-molybdopterin cofactor biosynthetic process [IEA |
| Molecular Function: |
catalytic activity
[IEA metal ion binding [IEA 4 iron, 4 sulfur cluster binding [IEA iron-sulfur cluster binding [IEA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSCRAGKKLF QWSRVKSSTE EIVKQLTVPL REHAEPILTL TPEQKREAVR LKIQEIEHTK 60
61 GQPPFFDMFM REHTYLRISL TEKCNFRCLY CMPAEGVPLK PKDKMLSNSE VLRLVKLFAA 120
121 HGVDKVRLTG GEPTIRKDIV HIVEGISSTP GIKEVGITTN GLVLQRFLPQ LRDAGLTKIN 180
181 ISIDSLDREK FAKMTRRDGF DKVWKAIELA RGYYPKVKLN VVVLKHQNEN EVVDFVNLTK 240
241 DRNLDVRFIE FMPFGGNEFK NDNFIGYREM LNLIVDKYGD GVIRLSDSPN DTTKAYKIDG 300
301 FQGQFGFITS MSDHFCNTCN RLRITADGNL KVCLHGNSEV SLRDRIRCGD SDEQLSEVIQ 360
361 KAVNNKKARH AVFRNGRSEE PAKSSNDSYR GLTPVTSASS ILVHLPSSSL YHSHLHSSRH 420
421 FFISQIRCFS TTYSVSSITH LLTHVDNNGN AKQVDVSQKD TSTRTAVARG TIILTAEISR 480
481 QISENTIKKG DVLTVAKIAS ILGAKQVANL IPLCHPIRLD FVDTVFNHDI ENSKLHCIST 540
541 ARCSGNTGVE MEALTACTIA LLTVYDMCKA ISQKMMLTNI YLVHKSGGKT TYTIDNENQI 600
601 |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
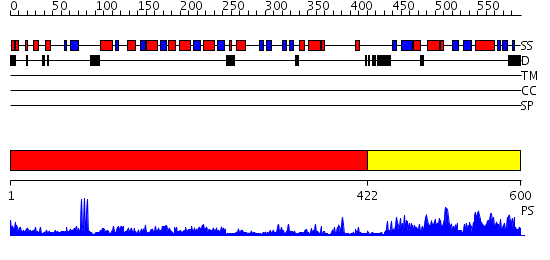
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..421] | 61.69897 | Structure of the S-adenosylmethionine dependent Enzyme MoaA | |
| 2 | View Details | [422..600] | 71.39794 | Molybdenum cofactor biosynthesis protein C, MoaC |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||
| 2 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.97 |
Source: Reynolds et al. (2008)