













| General Information: |
|
| Name(s) found: |
PH4alphaNE3-PA /
FBpp0085029
[FlyBase]
|
| Description(s) found:
Found 22 descriptions. SHOW ALL |
|
| Organism: | Drosophila melanogaster |
| Length: | 481 amino acids |
Gene Ontology: |
|
| Cellular Component: |
endoplasmic reticulum
[IEA]
procollagen-proline 4-dioxygenase complex [IC] |
| Biological Process: |
oxidation reduction
[IEA]
|
| Molecular Function: |
procollagen-proline 4-dioxygenase activity
[ISS]
L-ascorbic acid binding [IEA] oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen [IEA] iron ion binding [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MFCKVESTLN WESRQEFLDT ESAKMDLMQL DVELIDNLMN YAEKIDEKAS QLKRLAQELR 60
61 QPLHSAKGRE EEYLGNPLHS FPLIRHMYQD WRYLEEFMKK PVGEEEIQFL RRKLPELPWQ 120
121 VDTEEASVSI FRIAETYGMM PWDMANGLID NVRFNSTLPA LDCFEVAKMY FKWGYFKQAL 180
181 QWITISKARM KEEYSGVYEV LGMNRQDVAL LQARCLVELD RRDEAHEVLL DQPDLADNSI 240
241 SLLDQFKANP YEAIDSSPKL GEGYKRLCRS SFSPNPSKLH CRYNSTTSAF LILAPLKMEE 300
301 ISLEPHIVVY HDILPDKDIQ QLITLAEPLL KPTEMFDDNK NEARSSYRTP LGGPLLDSLT 360
361 QRMRDITGLQ IRQGNPINII KYGFGAPYTN YYDFFKKRNS ESKGFGDRMA TFMFYLNDAP 420
421 YGGATVFPRL NVKVPAERGK VLFWYNLNGD THDMEPTTMH AACPVFHGSK WVMTAWIHEY 480
481 D |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
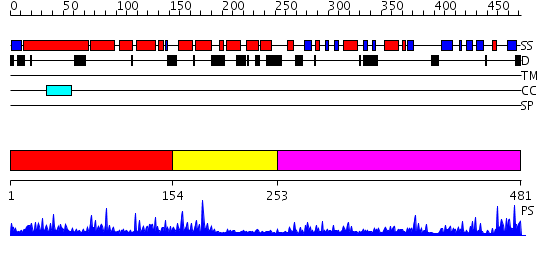
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..153] | 59.080922 | No description for PF08336.2 was found. No confident structure predictions are available. | |
| 2 | View Details | [154..252] | 13.221849 | Crystal structure of peptide-substrate-binding domain of human type I collagen prolyl 4-hydroxylase | |
| 3 | View Details | [253..481] | 4.95 | No description for 2hbtA was found. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||
| 1 | No functions predicted. | |||||||||||||||||||||||||||
| 2 |
|
|||||||||||||||||||||||||||
| 3 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)