













| General Information: |
|
| Name(s) found: |
SAC7 /
YDR389W
[SGD]
|
| Description(s) found:
Found 27 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 654 amino acids |
Gene Ontology: |
|
| Cellular Component: |
intracellular
[TAS |
| Biological Process: |
small GTPase mediated signal transduction
[IPI actin filament reorganization involved in cell cycle [IMP |
| Molecular Function: |
Rho GTPase activator activity
[IDA signal transducer activity [IPI |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MPNNTLKQGS KIENVSPSKG HVPSFWKQFI NNPKSMSSEN ITVPRSPTSL SRNAQPTTLK 60
61 RPPLSSRPYS YNTPTKDRKS FSKSAKQNNN NNNANSGTSP HAEFKNYRDM FLSNRNGFTG 120
121 RVFGVTLAES LSVASAEVIV QSELVSFGRI PIVVAKCGAY LKANGLETSG IFRIAGNGKR 180
181 VKALQYIFSS PPDYGTKFND WETYTVHDVA SLLRRYLNNL AEPLIPLSLY EQFRNPLRSR 240
241 PRILRHMLTH EVSHPNANKT NNVTVKSSRQ NYNDDGANDG DIEKEDAKDD EEKRRRKIRH 300
301 KRRLTRDIRA AIKEYEELFV TLSNDTKQLT IYLLDLLSLF ARQSQFNLMS GRNLAAIFQP 360
361 SILSHPQHDM DPKEYELSRL VVEFLIEYSY KLLPHLLKLA KREQQERLST ENKKNNGDKQ 420
421 KTDPIEIPKI TSSDSPPIVS SNKNPPAIDN NNKLDHTTLS PISTSIPENS SDLQTSKMLK 480
481 PPKQRRPHSK SFGSTPVPPD VIASNKRRTS LFPWLHKPGI LSDTGDNGDL TATEAEGDDY 540
541 EEENVDPYGQ SPSSVHSGSL PKQHYLPIPR MNRSLSGNST NSSFNTRPIS MILTSGNDNS 600
601 ADQLELLSNT HSNNERSNAL PLTEDDGDER NSRSRKRESW FQRLTSRSGS ANRA |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | Sample boi1 with gst from october 2005 | McCusker D, et al (2007) |
SHOWING SINGLE HITS. [ Hide Single Hits ]
| BAIT | PREY | HITS | PUBLICATION | |
| View Screen | 1 | Unpublished Fields Lab Data | ||
| View Screen | 1 | Unpublished Fields Lab Data |
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
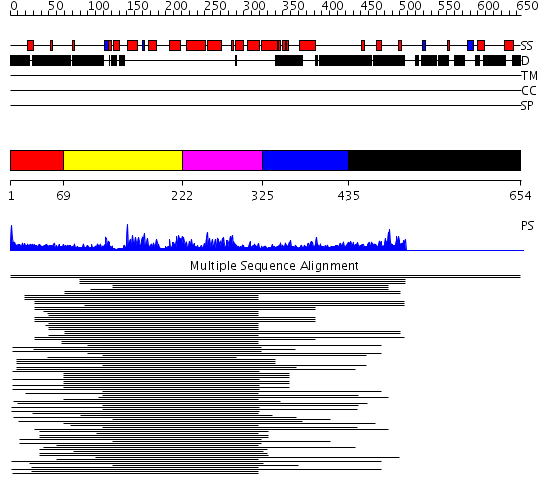
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..68] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [69..221] | 89.30103 | p50 RhoGAP domain | |
| 3 | View Details | [222..324] | 89.30103 | p50 RhoGAP domain | |
| 4 | View Details | [325..434] | 10.82 | p85 alpha subunit RhoGAP domain | |
| 5 | View Details | [435..654] | N/A | No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 5 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.93 |
Source: Reynolds et al. (2008)