













| General Information: |
|
| Name(s) found: |
gi|45440390
[NCBI NR]
gi|45435246 [NCBI NR] |
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Yersinia pestis biovar Microtus str. 91001 |
| Length: | 844 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSQDPFLERE AEKYESPIPS REFILAHLAK RETPASREEL ANELNLTGEE PLEALRRRLR 60
61 AMERDGQLVF TRRQCYALPE RLDLLRGTVI GHRDGYGFLR VEGSKDDLYL SAEQMKMAIH 120
121 GDVVLAQVVG ADRKGRREAR IVRVLVPKTS QIVGRYFTDA GTGFVVPDDS RLSFDILIPA 180
181 EATCGARMGF VVVVELTQRP TRRTKAVGKI VEVLGENMGT SMAVDIALRT HEIPHTWPPQ 240
241 VEKQVADLKE QVPEEAKKGR VDLRDLPLVT IDGEDARDFD DAVYCEKKRG GGWRLWVAIA 300
301 DVSYYVRPQT ALDDEARSRG NSVYFPSQVI PMLPEVLSNG LCSLNPQVDR LCMVCEMTIS 360
361 SQGKLSSYKF YEAVMSSHAR LTYTKVWRII DGEESLREQY KPLVPHLEEL HAMYKVLDQA 420
421 RAERGGIAFE TEEAKFIFNA ERRIERIEPT VRNDAHKLIE ECMILANIAA ARFVEKHEEP 480
481 ALFRVHDRPS DDHIVALRSV LNELGLTLGG GLKPQPKDYS VLMDEISDRP DHEMLQTMLL 540
541 RSMKQAVYDP ENRGHFGLAL ASYGHFTSPI RRYPDLAMHR AIKYQLAKEH GNLKDRWTPT 600
601 GGWHSDFEDM LQLGQHCSMT ERRADEATRD VADWLKCDFM LDQVGNVFTG IIASVTGFGF 660
661 FVRLDDLFID GLVHVSSLDN DYYRYDNIGQ RLIGESSGVV YRLGDTVEIR VDAVHMDERK 720
721 IDFALVSSTR KPRGEGKTAR DKAKNNGQRT LRSNSAGRGS SSGSSRRDGA PTAGGQKRPR 780
781 RAKKPVNFEP DSAFRPDAAA AKPDDKALAE KKAKAKAKRA SAKTKKIAAA TKAKRAKKKT 840
841 TSDQ |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
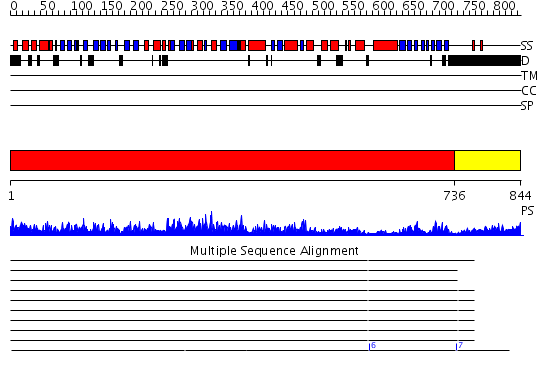
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..735] | 138.0 | No description for 2id0A was found. | |
| 2 | View Details | [736..844] | 8.522879 | Crystal structure of intact alpha subunit of aIF2 from Pyrococcus abyssi |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.97 |
Source: Reynolds et al. (2008)