













| General Information: |
|
| Name(s) found: |
TCPQ_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 12 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 549 amino acids |
Gene Ontology: |
|
| Cellular Component: |
membrane
[IDA]
|
| Biological Process: |
cellular protein metabolic process
[IEA]
protein folding [IEA] |
| Molecular Function: |
ATP binding
[IEA]
protein binding [IEA] unfolded protein binding [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MVGMSMQPYG IQSMLKEGYR HLSGLDEAVI KNIEACKELS TITRTSLGPN GMNKMVINHL 60
61 DKLFVTNDAA TIVNELEIQH PAAKLLVLAA KAQQEEIGDG ANLTISFAGE LLQNAEELIR 120
121 MGLHPSEIIS GYTKAVSKAV EILEQLVETG SETMDVRNKD EVISRMRAAV ASKQFGQEEI 180
181 ICSLVTDACI QVCPKNPTNF NVDNVRVSKL LGGGLHNSCI VRGMVLKSDA VGSIKRMEKA 240
241 KVAVFAGGVD TTATETKGTV LIHSAEQLEN YAKTEEAKVE ELIKAVAESG AKVIVSGGSI 300
301 GEMALHFCER YKIMVLKISS KFELRRFCRT AGAVAHLKLS RPSPEDLGYV DSISVEEIGG 360
361 VTVTIARNEE GGNSISTVVL RGSTDSILDD LERAVDDGVN TYKAMCRDSR IVPGAAATEI 420
421 ELAQRLKEYA NAEIGLDKYA ITKYAESFEF VPKTLADNAG LNAMEIIAAL YTGHGSGNTK 480
481 LGIDLEEGAC KDVSETKVWD LFATKLFALK YASDAACTVL RVDQIIMAKP AGGPRRDAAQ 540
541 AAGAGAEED |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
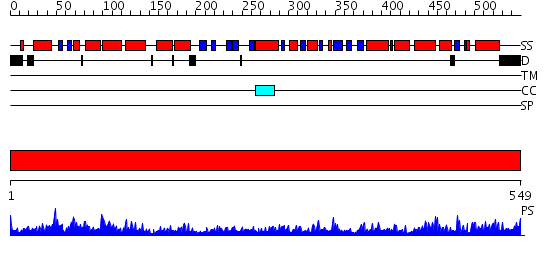
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..549] | 147.0 | Crystal structure of the chaperonin from Thermococcus strain KS-1 (nucleotide-free form of single mutant) |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)