













| General Information: |
|
| Name(s) found: |
GDPD2_ARATH
[Swiss-Prot]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Arabidopsis thaliana |
| Length: | 374 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
glycerol metabolic process
[IEA][ISS]
lipid metabolic process [IEA] |
| Molecular Function: |
glycerophosphodiester phosphodiesterase activity
[IEA][ISS]
phosphoric diester hydrolase activity [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MALRTVLVSD VPSLPDSVYG LSEGLELSKP TSFRLPGFSV IGHRGIGMNV LQSSDRRARG 60
61 VKENSILSFN SAAKYPIDFI EFDVQVTKDD CPVIFHDDFI YSEENGIVNE SRVTDLSLSE 120
121 FLLYGPQKET EKIGKTLMRK SKEGKVLKWD VDLDDSLCTL QEAFEQVEQT LGFNIELKFD 180
181 DQTVYEREFL VHILRSVLQV VSNYAKDRPV IFSSFQPDAA KLVRELQSTY PVFFLTDAGN 240
241 EIHNDERRNS LEEAIQVCLE GGLQGIVSEV KGVFRNPAAI SKIKESNLSL LTYGKLNNVG 300
301 EAVYMQYVMG IDGVIVDFVE EIIESTTRMM IRPPPSSSPL PSPSKDDDVA ITRPEFSQKE 360
361 ISFLLKLLSQ LIQH |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
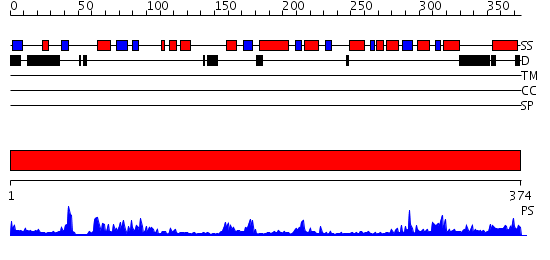
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..374] | 53.39794 | Crystal structure of periplasmic glycerophosphodiester phosphodiesterase from Escherichia coli |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.91 |
Source: Reynolds et al. (2008)