













| General Information: |
|
| Name(s) found: |
CCA1_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 16 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 608 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IDA]
|
| Biological Process: |
response to gibberellin stimulus
[IEP]
response to abscisic acid stimulus [IEP] response to salt stress [IEP] long-day photoperiodism, flowering [IGI] response to jasmonic acid stimulus [IEP] response to cadmium ion [IEP] response to salicylic acid stimulus [IEP] circadian rhythm [IMP][TAS] response to organic nitrogen [IEP] response to ethylene stimulus [IEP] response to auxin stimulus [IEP] |
| Molecular Function: |
DNA binding
[IDA]
transcription factor activity [ISS] transcription activator activity [IMP] transcription repressor activity [IEP] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 METNSSGEDL VIKTRKPYTI TKQRERWTEE EHNRFIEALR LYGRAWQKIE EHVATKTAVQ 60
61 IRSHAQKFFS KVEKEAEAKG VAMGQALDIA IPPPRPKRKP NNPYPRKTGS GTILMSKTGV 120
121 NDGKESLGSE KVSHPEMANE DRQQSKPEEK TLQEDNCSDC FTHQYLSAAS SMNKSCIETS 180
181 NASTFREFLP SREEGSQNNR VRKESNSDLN AKSLENGNEQ GPQTYPMHIP VLVPLGSSIT 240
241 SSLSHPPSEP DSHPHTVAGD YQSFPNHIMS TLLQTPALYT AATFASSFWP PDSSGGSPVP 300
301 GNSPPNLAAM AAATVAAASA WWAANGLLPL CAPLSSGGFT SHPPSTFGPS CDVEYTKAST 360
361 LQHGSVQSRE QEHSEASKAR SSLDSEDVEN KSKPVCHEQP SATPESDAKG SDGAGDRKQV 420
421 DRSSCGSNTP SSSDDVEADA SERQEDGTNG EVKETNEDTN KPQTSESNAR RSRISSNITD 480
481 PWKSVSDEGR IAFQALFSRE VLPQSFTYRE EHREEEQQQQ EQRYPMALDL NFTAQLTPVD 540
541 DQEEKRNTGF LGIGLDASKL MSRGRTGFKP YKRCSMEAKE SRILNNNPII HVEQKDPKRM 600
601 RLETQAST |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
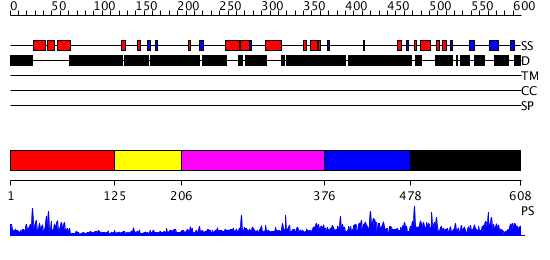
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..124] | 12.69897 | No description for 2v1dB was found. | |
| 2 | View Details | [125..205] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [206..375] | 1.260988 | View MSA. No confident structure predictions are available. | |
| 4 | View Details | [376..477] | 2.086983 | View MSA. No confident structure predictions are available. | |
| 5 | View Details | [478..608] | 3.017975 | View MSA. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.64 |
Source: Reynolds et al. (2008)