













| General Information: |
|
| Name(s) found: |
FBpp0303400
[FlyBase]
Dhap-at-PA / FBpp0084271 [FlyBase] |
| Description(s) found:
Found 26 descriptions. SHOW ALL |
|
| Organism: | Drosophila melanogaster |
| Length: | 724 amino acids |
Gene Ontology: |
|
| Cellular Component: |
peroxisome
[ISS][NAS]
mitochondrion [ISS] |
| Biological Process: |
metabolic process
[IEA]
|
| Molecular Function: |
glycerone-phosphate O-acyltransferase activity
[ISS][NAS]
glycerol-3-phosphate O-acyltransferase activity [ISS] acyltransferase activity [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MDRNPVRQST SSGTVEGESS EQEEQKTPSA AEYMRNFKNI IAPGNEASMT REFNPQVAYE 60
61 FEKYLNPQKL KQHVLRSEKL RSILEHYAKE SGTPLKQMER QARAIIDEIG LDRNMAIIRW 120
121 CGIAITAIGK RICDGFYVNS ASMANVRKDM GKCPVLYLPS HRSYMDFILM SYICYYYDIE 180
181 IPGIAAGMDF HSMFGMGTML RKTGAFFMRR SFSNDELYWD IFREYMYALV ANYHIGVEFF 240
241 IEGTRSRNFK ALVPKIGLLS MALLPYFTGE VPDVMIVPVS VAYERVLEEQ LFVYELLGVP 300
301 KPKESTKGFF KALKIIDERF GKMFLDFGEP ISVKEFFGHD SAQRMQRAGV GGHLQKLNRQ 360
361 EVELVKQLAN EIIYQQQRRI VISTFNLLSL YYASQLYAQR SVTLDELARG VLHLKRIFEQ 420
421 LGAHVSTTPS SIKADVIDAV EIHSNILHFA TGRLQFTPMA AKQLAEQIDV KRLKAHALCP 480
481 QTMAVAVPML ALQLYINPCM FWLARPAYLL LAALKEQKNQ QKQSTDISYD ASLCALHAHV 540
541 TTMDALFQHE FIIESNREAA EFETHLQLLL DERVVEVETS GRINVQDNEC SHVILAALAP 600
601 FLCLYYQLVV TLRKIPLELE FSNKELLVRV QQHVEQLLQQ PGASASHVHP YCLALDNLNI 660
661 AIYALIQRGY LVKSRDSGQM KIASTPGKCL RELETQLLEY CQLMPFAQYS TAANQQAQFH 720
721 IAKL |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
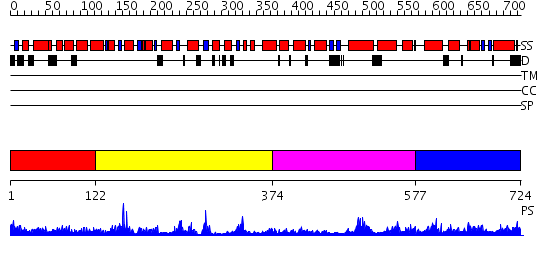
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..121] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [122..373] | 8.522879 | The 1.55 A Crystal Structure of Glycerol-3-Phosphate Acyltransferase | |
| 3 | View Details | [374..576] | 1.166987 | View MSA. No confident structure predictions are available. | |
| 4 | View Details | [577..724] | 2.10598 | View MSA. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.71 |
Source: Reynolds et al. (2008)