













| General Information: |
|
| Name(s) found: |
gi|24375413
[NCBI NR]
gi|24350251 [NCBI NR] |
| Description(s) found:
Found 5 descriptions. SHOW ALL |
|
| Organism: | Shewanella oneidensis MR-1 |
| Length: | 359 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
biotin biosynthetic process
[ISS |
| Molecular Function: |
biotin synthase activity
[ISS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MITRPSPPAP VTQPTSVKPT LLNTVFSYAE ILSLLQGQDD EWLFSRAKLA TELEFNQQVY 60
61 LRGIVEFSNH CRNHCHYCGL RTENRQVTRY RLSNEEILNA VDSIAELGLG TVVLQSGDDF 120
121 NYSGNRISTL ITEIKRHHNL AITLSLGDRK HQELEKWREA GADRYLLKME TFDRALFAQC 180
181 RPKANFDERI ARLNYLKSLG YQTGSGIIVD LPGMTDAILA RDIQHLSELQ LDMLACGPFI 240
241 AHHQTPFTTS PNGSALKSHR VSAILRLMNP GANIPATSSL DALDKGAREQ ALKRGCNVIM 300
301 PSFTPTKVSG DYSIYPGKNQ QQHPAAERLN QVCQQIQRHG LIPSFSRGDS KRTQYVSRH |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
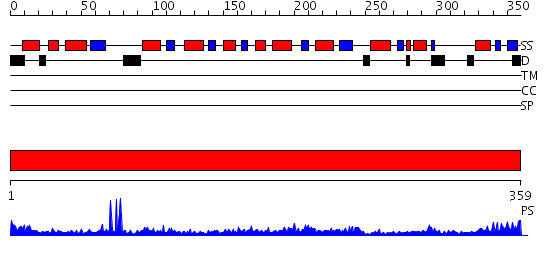
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..359] | 57.69897 | The Crystal Structure of Biotin Synthase, an S-Adenosylmethionine-Dependent Radical Enzyme |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.97 |
Source: Reynolds et al. (2008)