













| General Information: |
|
| Name(s) found: |
SPA2_YEAST
[Swiss-Prot]
|
| Description(s) found: |
|
| Organism: | Saccharomyces cerevisiae S288c |
| Length: | 1466 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MGTSSEVSLA HHRDIFHYYV SLKTFFEVTG ENRDRSNSTR AQKARAKLLK LSSSQFYELS 60
61 TDVSDELQRR IGEDANQPDY LLPKANFHMK RNQARQKLAN LSQTRFNDLL DDILFEIKRR 120
121 GFDKDLDAPR PPLPQPMKQE VSKDSDDTAR TSTNSSSVTQ VAPNVSVQPS LVIPKMASID 180
181 WSSEEEEEEQ VKEKPNEPEG KQTSMDEKKE AKPALNPIVT DSDLPDSQVL ARDITSMART 240
241 PTTTHKNYWD VNDSPIIKVD KDIDNEKGPE QLKSPEVQRA ENNNPNSEME DKVKELTDLN 300
301 SDLHLQIEDL NAKLASLTSE KEKEKKEEKE EKEKEKNLKI NYTIDESFQK ELLSLNSQIG 360
361 ELSIENENLK QKISEFELHQ KKNDNHNDLK ITDGFISKYS SADGLIPAQY ILNANNLIIQ 420
421 FTTRLSAVPI GDSTAISHQI GEELFQILSQ LSNLISQLLL SADLLQYKDQ VILLKASLSH 480
481 AITSIRYFSV YGPVLIPKIT VQAAVSEVCF AMCNLIDSAK IKSDSNGEST TSNEGNRQVL 540
541 EYSSPTATTP MTPTFPSTSG INMKKGFINP RKPASFLNDV EEEESPVKPL KITQKAINSP 600
601 IIRPSSSNGV PTTSRKPSGT GLFSLMIDSS IAKNSSHKED NDKYVSPIKA VTSASNSASS 660
661 NISEIPKLTL PPQAKIGTVI PPSENQVPNI KIENTEEDNK RSDITNEISV KPTSSIADKL 720
721 KQFEQSSEKK SSPKENPIAK EEMDSKPKLS NKFITSMNDV STDDSSSDGN ENDDADDDDD 780
781 FTYMALKQTM KREGSKIEKN NDSKLPANIV ELDLHESPES VKIESPESIK EITSSEMSSE 840
841 MPSSSLPKRL VEDVEPSEMP EKGASVESVR KKNFQEPLGN VESPDMTQKV KSLGMTGKAV 900
901 GPESDSRVES PGMTGQIKSL NMAGKVVGPE ADSRVESPGM KEQIKSLGMT GKITAQESIK 960
961 SPEAARKLAS SGEVDKIESP RMVRESESLE AVGNTIPSNM TVKMESPNLK GNTVSEPQEI1020
1021 RRDIASSEPI ENVDPPKVLK KIVFPKAVNR TGSPKSVEKT PSSATLKKSG LPEPNSQIVS1080
1081 PELAKNSPLA PIKKNVELRE TNKPHTETIT SVEPTNKDAN TSWRDADLNR TIKREEEDED1140
1141 FDRVNHNIQI TGAYTKTGKI DYHKIPVDRK AKSEAEVHTS EEDIDESNNV NGKRADAQIH1200
1201 ITERKHAFVN PTENSQVKKT SHSPFLNSKP VQYENSESNG GINNHIKIKN TGETTAHDEK1260
1261 HYSDDDDSSY QFVPMKHEEQ EQEQNRSEEE ESEDDDEEEE DSDFDVDTFD IENPDNTLSE1320
1321 LLLYLEHQTM DVISTIQSLL TSIKKPQVTK GNLRGESNAI NQVIGQMVDA TSISMEQSRN1380
1381 ANLKKHGDWV VQSLRDCSRR MTILCQLTGD GILAKEKSDQ DYADKNFKQR LAGIAFDVAK1440
1441 CTKELVKTVE EASLKDEINY LNSKLK |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
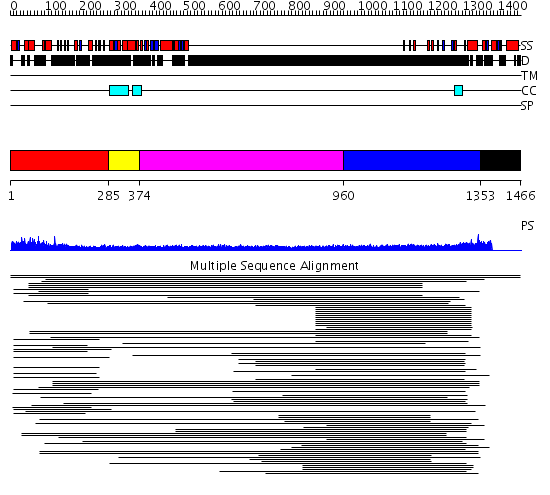
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..284] | 7.060967 | View MSA. No confident structure predictions are available. | |
| 2 | View Details | [285..373] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [374..959] | 1.163997 | View MSA. No confident structure predictions are available. | |
| 4 | View Details | [960..1352] | 24.221997 | View MSA. No confident structure predictions are available. | |
| 5 | View Details | [1353..1466] | N/A | Confident ab initio structure predictions are available. | |
| 6 | View Details | [825..947] | N/A | No confident structure predictions are available. | |
| 7 | View Details | [948..1100] | N/A | No confident structure predictions are available. | |
| 8 | View Details | [1101..1353] | 2.403996 | View MSA. No confident structure predictions are available. | |
| 9 | View Details | [1354..1466] | N/A | Confident ab initio structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)