GO TERM SUMMARY
|
| Name: |
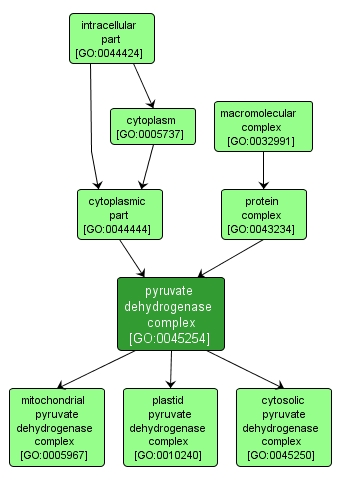
pyruvate dehydrogenase complex |
| Acc: |
GO:0045254 |
| Aspect: |
Cellular Component |
| Desc: |
Complex that carries out the oxidative decarboxylation of pyruvate to form acetyl-CoA; comprises subunits possessing three catalytic activities: pyruvate dehydrogenase (E1), dihydrolipoamide S-acetyltransferase (E2), and dihydrolipoamide dehydrogenase (E3). |
Synonyms:
- dihydrolipoyl dehydrogenase complex
- pyruvate dehydrogenase complex (lipoamide)
- GO:0009364
|
|

|
INTERACTIVE GO GRAPH
|