













| General Information: |
|
| Name(s) found: |
gi|31455207
[NCBI NR]
gi|20986495 [NCBI NR] gi|119573439 [NCBI NR] |
| Description(s) found:
Found 19 descriptions. SHOW ALL |
|
| Organism: | Homo sapiens |
| Length: | 684 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MKKPSAKQQK EVEKVKPQCK EVHQTLILDP AQRKRLQQQM QQHVQLLTQI HLLATCNPNL 60
61 NPEASSTRIC LKELGTFAQS SIALHHQYNP KFQTLFQPCN LMGAMQLIED FSTHVSIDCS 120
121 PHKTVKKTAN EFPCLPKQVA WILATSKVFM YPELLPVCSL KAKNPQDKIL FTKAEDNLLA 180
181 LGLKHFEGTE FLNPLISKYL LTCKTARQLT VRIKNLNMNR APDNIIKFYK KTKQLPVLGK 240
241 CCEEIQPHQW KPPIEREEHR LPFWLKASLP SIQEELRHMA DGAREVGNMT GTTEINSDQG 300
301 LEKDNSELGS ETRYPLLLPK GVVLKLKPVA DRFPKKAWRQ KRSSVLKPLL IQPSPSLQPS 360
361 FNPGKTPAQS THSEAPPSKM VLRIPHPIQP ATVLQTVPGV PPLGVSGGES FESPAALPAM 420
421 PPEARTSFPL SESQTLLSSA PVPKVMMPSP ASSMFRKPYV RRRPSKRRGA RAFRCIKPAP 480
481 VIHPASVIFT VPATTVKIVS LGGGCNMIQP VNAAVAQSPQ TIPIATLLVN PTSFPCPLNQ 540
541 PLVASSVSPL IVSGNSVNLP IPSTPEDKAH MNVDIACAVA DGENAFQGLE PKLEPQELSP 600
601 LSATVFPKVE HSPGPPPVDK QCQEGLSENS AYRWTVVKTE EGRQALEPLP QGIQESLNNS 660
661 SPGDLEEVVK MEPEDATEEI SGFL |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | No Comments | Cheeseman, I.M., et al. (2004) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
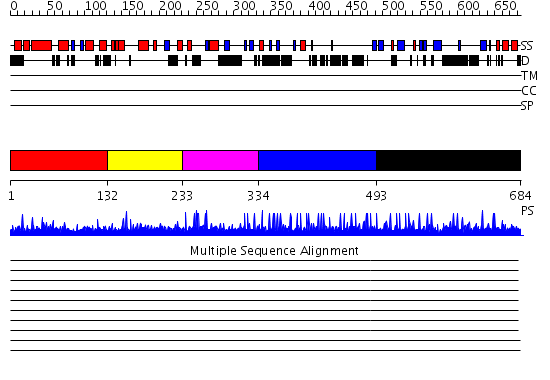
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..131] | 1.019982 | View MSA. No confident structure predictions are available. | |
| 2 | View Details | [132..232] | 2.021993 | View MSA. No confident structure predictions are available. | |
| 3 | View Details | [233..333] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [334..492] | 1.019996 | View MSA. No confident structure predictions are available. | |
| 5 | View Details | [493..684] | 2.015908 | View MSA. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.95 |
Source: Reynolds et al. (2008)