













| General Information: |
|
| Name(s) found: |
YJEK_ECOLI
[Swiss-Prot]
|
| Description(s) found:
Found 8 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli |
| Length: | 342 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAHIVTLNTP SREDWLTQLA DVVTDPDELL RLLNIDAEEK LLAGRSAKKL FALRVPRSFI 60
61 DRMEKGNPDD PLLRQVLTSQ DEFVIAPGFS TDPLEEQHSV VPGLLHKYHN RALLLVKGGC 120
121 AVNCRYCFRR HFPYAENQGN KRNWQTALEY VAAHPELDEM IFSGGDPLMA KDHELDWLLT 180
181 QLEAIPHIKR LRIHSRLPIV IPARITEALV ECFARSTLQI LLVNHINHAN EVDETFRQAM 240
241 AKLRRVGVTL LNQSVLLRDV NDNAQTLANL SNALFDAGVM PYYLHVLDKV QGAAHFMVSD 300
301 DEARQIMREL LTLVSGYLVP KLAREIGGEP SKTPLDLQLR QQ |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
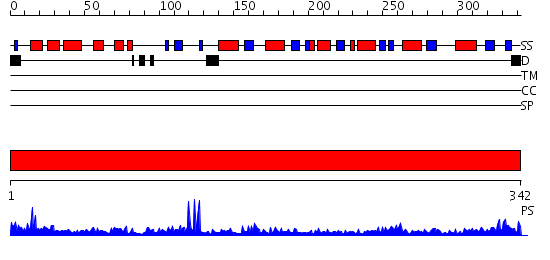
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..342] | 90.09691 | 2.1 Angstrom X-ray crystal structure of lysine-2,3-aminomutase from Clostridium subterminale SB4, with Michaelis analog (L-alpha-lysine external aldimine form of pyridoxal-5'-phosphate). |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)