













| General Information: |
|
| Name(s) found: |
YGEV_ECOLI
[Swiss-Prot]
|
| Description(s) found:
Found 9 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli |
| Length: | 592 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MELATTQSVL MQIQPTIQRF ARMLASVLQL EVEIVDENLC RVAGTGAYGK FLGRQLSGNS 60
61 RLLRHVLETK TEKVVTQSRF DPLCEGCDSK ENCREKAFLG TPVILQDRCV GVISLIAVTH 120
121 EQQEHISDNL REFSDYVRHI STIFVSKLLE DQGPGDNISK IFATMIDNMD QGVLVVDDEN 180
181 RVQFVNQTAL KTLGVVQNNI IGKPIRFRPL TFESNFTHGH MQHIVSWDDK SELIIGQLHN 240
241 IQGRQLFLMA FHQSHTSFSV ANAPDEPHIE QLVGECRVMR QLKRLISRIA PSPSSVMVVG 300
301 ESGTGKEVVA RAIHKLSGRR NKPFIAINCA AIPEQLLESE LFGYVKGAFT GASANGKTGL 360
361 IQAANTGTLF LDEIGDMPLM LQAKLLRAIE AREILPIGAS SPIQVDIRII SATNQNLAQF 420
421 IAEGKFREDL FYRLNVIPIT LPPLRERQED IELLVHYFLH LHTRRLGSVY PGIAPDVVEI 480
481 LRKHRWPGNL RELSNLMEYL VNVVPSGEVI DSTLLPPNLL NNGTTEQSDV TEVSEAHLSL 540
541 DDAGGTALEE MEKQMIREAL SRHNSKKQVA DELGIGIATL YRKIKKYELL NT |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
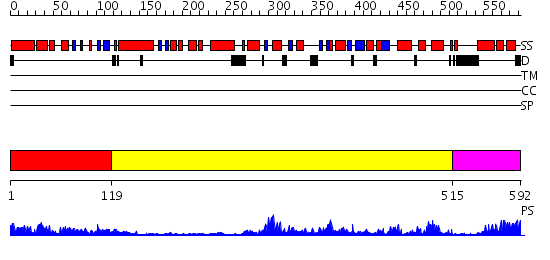
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..118] | 22.0 | ClpB | |
| 2 | View Details | [119..514] | 83.522879 | Crystal structure of sigm54 activator (AAA+ ATPase) in the inactive state | |
| 3 | View Details | [515..592] | 80.30103 | Crystal structure of a sigma54-activator suggests the mechanism for the conformational switch necessary for sigma54 binding |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||||||||||||||||||||||||||
| 2 |
|
|||||||||||||||||||||||||||||||||||||||||||||
| 3 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)