













| General Information: |
|
| Name(s) found: |
HSCC_ECOLI
[Swiss-Prot]
|
| Description(s) found:
Found 9 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli |
| Length: | 556 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MDNAELAIGI DLGTTNSLIA VWKDGAAQLI PNKFGEYLTP SIISMDENNH ILVGKPAVSR 60
61 RTSHPDKTAA LFKRAMGSNT NWRLGSDTFN APELSSLVLR SLKEDAEEFL QRPIKDVVIS 120
121 VPAYFSDEQR KHTRLAAELA GLNAVRLINE PTAAAMAYGL HTQQNTRSLV FDLGGGTFDV 180
181 TVLEYATPVI EVHASAGDNF LGGEDFTHML VDEVLKRADV ARTTLNESEL AALYACVEAA 240
241 KCSNQSPLHI RWQYQEETRE CEFYENELED LWLPLLNRLR VPIEQALRDA RLKPSQIDSL 300
301 VLVGGASQMP LVQRIAVRLF GKLPYQSYDP STIVALGAAI QAACRLRSED IEEVILTDIC 360
361 PYSLGVEVNR QGVSGIFSPI IERNTTVPVS RVETYSTMHP EQDSITVNVY QGENHKVKNN 420
421 ILVESFDVPL KKTGAYQSID IRFSYDINGL LEVDVLLEDG SVKSRVINHS PVTLSAQQIE 480
481 ESRTRLSALK IYPRDMLINR TFKAKLEELW ARALGDEREE IGRVITDFDA ALQSNDMARV 540
541 DEVRRRASDY LAIEIP |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
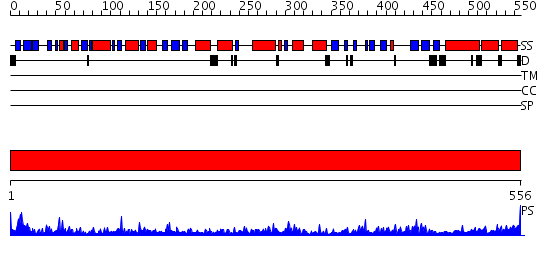
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..556] | 156.0 | crystal structure of bovine hsc70(aa1-554)E213A/D214A mutant |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.98 |
Source: Reynolds et al. (2008)