













| General Information: |
|
| Name(s) found: |
MSRB4_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 10 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 139 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytosol
[ISS]
|
| Biological Process: |
biological_process
[ND]
|
| Molecular Function: |
peptide-methionine-(S)-S-oxide reductase activity
[ISS]
|
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
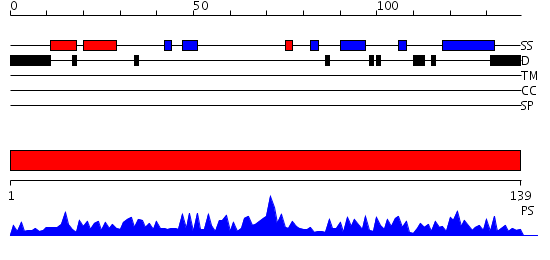
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..139] | 53.69897 | C-terminal MsrB domain of peptide methionine sulfoxide reductase PilB |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)