













| General Information: |
|
| Name(s) found: |
ACLB2_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 15 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 608 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytosol
[IDA]
citrate lyase complex [IDA] plasma membrane [IDA] |
| Biological Process: |
acetyl-CoA biosynthetic process
[TAS]
|
| Molecular Function: |
ATP citrate synthase activity
[IDA]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MATGQLFSRT TQALFYNYKQ LPVQRMLDFD FLCGRETPSV AGIINPGSEG FQKLFFGQEE 60
61 IAIPVHAAIE AACAAHPTAD VFINFASFRS AAASSMAALK QPTIKVVAII AEGVPESDTK 120
121 QLIAYARANN KVVIGPATVG GIQAGAFKIG DTAGTIDNII QCKLYRPGSV GFVSKSGGMS 180
181 NEMYNTVARV TDGIYEGIAI GGDVFPGSTL SDHILRFNNI PQIKMMVVLG ELGGRDEYSL 240
241 VEALKEGKVN KPVVAWVSGT CARLFKSEVQ FGHAGAKSGG EMESAQAKNQ ALIDAGAIVP 300
301 TSFEALESAI KETFEKLVEE GKVSPIKEVI PPQIPEDLNS AIKSGKVRAP THIISTISDD 360
361 RGEEPCYAGV PMSSIIEQGY GVGDVISLLW FKRSLPRYCT KFIEICIMLC ADHGPCVSGA 420
421 HNTIVTARAG KDLVSSLVSG LLTIGPRFGG AIDDAARYFK DACDRNLTPY EFVEGMKKKG 480
481 IRVPGIGHRI KSRDNRDKRV ELLQKFARSN FPSVKYMEYA VTVETYTLSK ANNLVLNVDG 540
541 AIGSLFLDLL AGSGMFTKQE IDEIVQIGYL NGLFVLARSI GLIGHTFDQK RLKQPLYRHP 600
601 WEDVLYTK |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
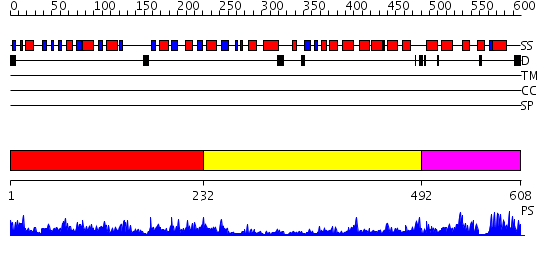
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..231] | 82.154902 | Succinyl-CoA synthetase, alpha-chain, N-terminal (CoA-binding) domain; Succinyl-CoA synthetase, alpha-chain, C-terminal domain | |
| 2 | View Details | [232..491] | 1.67 | No description for 2ifcA was found. | |
| 3 | View Details | [492..608] | 92.30103 | Citrate synthase |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||
| 2 | No functions predicted. | ||||||||||||||||||||||||||||||
| 3 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.83 |
Source: Reynolds et al. (2008)