













| General Information: |
|
| Name(s) found: |
gi|9294015
[NCBI NR]
gi|15231330 [NCBI NR] |
| Description(s) found:
Found 10 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 753 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cellular_component
[ND]
|
| Biological Process: |
protein amino acid phosphorylation
[IEA]
|
| Molecular Function: |
ATP binding
[IEA]
protein tyrosine kinase activity [IEA] protein kinase activity [IEA] kinase activity [ISS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSRLQKRGKQ EKPVVSDGAQ KVIVAVKASR EIPKTALIWA LTHVVQPGDC ITLIVVVPSH 60
61 NSGRKLWGFT KSFPMFAGDC ASGHRKSHSE ALPEIKSDLT DTCSQMILQL HDVYDPNKIN 120
121 VKIKIVSGSP CGAVAAESKK AQANWVVMDK HLKQEEKRCM DELQCNIVVM KRSQAKVLRL 180
181 NLVGSPKKDA GKECPLPSGP EAASEKHSKN TKGLLDADRG LPVTPTSSPE LGTPFTSTEA 240
241 GTSSVSSSDL GTSPFFTLGM NGYMKKDGAL VIKENDGLDD SGSETESENQ SLASTSMRFQ 300
301 PWISEYIGTH RHSSQEAEES LLWKNDDRAQ ISTTKALLEK FSKLDVEVGL SSSRRMDLEF 360
361 SGNVRDAISL SRSAPPGPPP LCSICQHKAP VFGKPPRLFT YAELELATGG FSQANFLAEG 420
421 GYGSVHRGVL PEGQVVAVKQ HKLASSQGDV EFCSEVEVLS CAQHRNVVML IGFCIEDSRR 480
481 LLVYEYICNG SLDSHLYGRQ KETLEWPARQ KIAVGAARGL RYLHEECRVG CIVHRDMRPN 540
541 NILITHDNEP LVGDFGLARW QPDGEMGVDT RVIGTFGYLA PEYAQSGQIT EKADVYSFGV 600
601 VLVELVTGRK AIDITRPKGQ QCLTEWARPL LEEYAIDELI DPRLGNRFVE SEVICMLHAA 660
661 SLCIRRDPHL RPRMSQVLRI LEGDMIMDGN YASTPGSEAG NRSGRFWADH YSGQLTNDGS 720
721 DRFSERLSVE TPRLALRERE RSQRFELNHN KQY |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
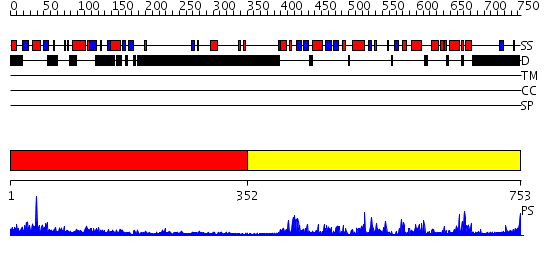
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..351] | 32.69897 | Crystal Structure of the 3 Ig form of FGFR3c in complex with FGF1 | |
| 2 | View Details | [352..753] | 71.0 | No description for 2qkwB was found. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.92 |
Source: Reynolds et al. (2008)