













| General Information: |
|
| Name(s) found: |
tba-4 /
CE18680
[WormBase]
|
| Description(s) found:
Found 20 descriptions. SHOW ALL |
|
| Organism: | Caenorhabditis elegans |
| Length: | 448 amino acids |
Gene Ontology: |
|
| Cellular Component: |
protein complex
[IEA microtubule [IEA |
| Biological Process: |
protein polymerization
[IEA pronuclear migration [IMP locomotory behavior [IMP morphogenesis of an epithelium [IMP nematode larval development [IMP embryonic development ending in birth or egg hatching [IMP positive regulation of growth rate [IMP gamete generation [IMP growth [IMP microtubule-based process [IEA |
| Molecular Function: |
GTP binding
[IEA GTPase activity [IEA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MREVISIHVG QAGVQIGNAC WELYCLEHGI QPDGTMPSEQ QNEGGSFTTF FSDTGNGRYV 60
61 PRSIFVDLEP TVVDEIRTGN YKKLFHPEQM ITGKEDAANN YARGHYTVGK ELIDTTLDRI 120
121 RRLADNCSGL QGFFVFHSFG GGTGSGFTSL LMERLSVDYG KKSKLEFSIY PAPQVSTAVV 180
181 EPYNSILTTH TTLEHSDCAF MVDNEAIYDI CKRNLDVDRP SYTNLNRIIS QVVSSITASL 240
241 RFDGALNVDL NEFQTNLVPY PRIHFPLASY TPLISAEKAY HEALSVNDIT NSCFEPANQM 300
301 VKCDPRNGKY MAVCLLYRGD VVPKDVNTAI TAIKTKRSVQ FVDWCPTGFK VGINYQPPTV 360
361 VPGGDLAKVP RAVCMLSNTT AIAEAWSRLD YKFDLMYAKR AFVHWYVGEG MEEGEFTEAR 420
421 EDLAALEKDY EEVGADSNEG LEEDGEEY |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | No Comments | Cheeseman, I.M., et al. (2004) | |
| View Run | hcp1 IP from december2004 | Cheeseman, I.M., et al. (2005) | |
| View Run | hcp2 ip from december | Cheeseman, I.M., et al. (2005) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
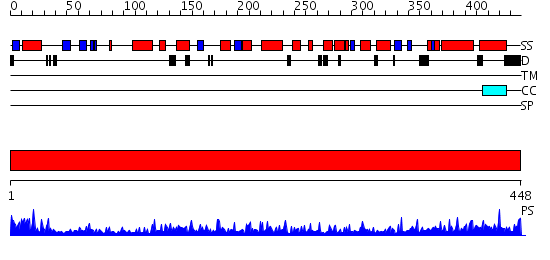
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..448] | 176.0 | Tubulin-colchicine-vinblastine: stathmin-like domain complex |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.78 |
Source: Reynolds et al. (2008)