













| General Information: |
|
| Name(s) found: |
lin-66 /
CE15557
[WormBase]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Caenorhabditis elegans |
| Length: | 627 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
hermaphrodite genitalia development
[IMP]
locomotion [IMP] nematode larval development [IMP] reproduction [IMP] positive regulation of locomotion [IMP] |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSYEMNSLFS SNQQPLGGGP PPQQQSQQQQ QPPQSLWSLQ QFSQMGAPDN FGNHAVPFMP 60
61 TLGMLNQYQN PHPLQMVPPQ PLVTPSQVLP PSMAEQQRAD TIHGFGVLTW LSAKAGLITC 120
121 KDKMIISFQL KDFCDQMLND LTSVLRVGFT LSFHAALNET SEYTATIVSP IYGPDADVLF 180
181 ANSQEVDLEA ANPTPPNSKD AYSPALEQKA IPALLAIFQR HGLQQIQLSR SFSLHSQMSN 240
241 CGDDELFRYV GTSSLKRRQF VERRTHLFRL QNDDSIILQY PAVYQAVYQL ASYLLRRGGA 300
301 TSIQSLFDYY GSPDIMPEVR SHTGEGRQDF LNLLTAHNWV FALFPNRTYV SVRRNLPNYD 360
361 YVGFIKQHFQ EMDITRSQQM AYGQPPRGIQ RTMSAQMGSS FATGRPQNLL MQHQHAMTPA 420
421 ATRSGSLWDS QSSLSQSGNS GDQWSPLFSP STWSNNTNAN PRLTSLTPLS SMNETLLSGV 480
481 NYNSLLAMPP SAKNERSIGV QADDFLDRMS YNPIGRTGCT CQCQCGRGGA AIIGNTRGSS 540
541 SRASGGSASP PSADGASQID RFSPSNHAGS IGDRTLTAAA LGVGLVDQQP AAPNHDGSAN 600
601 QPQRYYDPFG TADLLNGTRL NSLRIGN |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | hcp1 IP from december2004 | Cheeseman, I.M., et al. (2005) |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
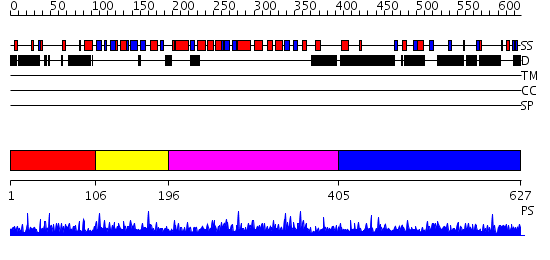
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..105] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [106..195] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [196..404] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [405..627] | 1.004947 | View MSA. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.98 |
Source: Reynolds et al. (2008)