













| General Information: |
|
| Name(s) found: |
gi|19113470
[NCBI NR]
|
| Description(s) found:
Found 2 descriptions. SHOW ALL |
|
| Organism: | Schizosaccharomyces pombe 972h- |
| Length: | 827 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSSLASLIKT KLKLPDSSKN LNKLRWKFRL LESQFALYSL PCQLRFVHYN NLEDPNSFNG 60
61 LLYLFTDYIA FQGDDESNQF CMPYTIIRKV SRVKSNDLEQ LLSVSTSNGY EYRISLQVSE 120
121 TTAAHFCQLL REELVSHKAD MQRSSEFSKQ FFSERLCKPN VPEPETKDSY GFGARYGYPT 180
181 DPRISRERAK LRMWKEYFLL YGANLSLIRV SLFSKLVRIE LPNKLRGEIW ELTSGSMYFR 240
241 LENSDEYDHL LKVYSGQTSF SLEEIEKDLG RSLPEYPAYQ NEEGINALRN VLVAFSWKNQ 300
301 EVGYCQAMNI VAAALLIHCT EEQTFFLMHK ICEDYIPGYY SKTMYGTLID QQVYESLVQR 360
361 SMPNLHAHFV SKDIQLSIIS LPWFLSLFLC TMPLPYAFRL LDFFFLEGPR VLFQIGMAIL 420
421 YDNEAEIMKA TEDTMLISIL KNYFSSLGDK AYKDATDKRV ASITKFQLLL VTAFKKFSHI 480
481 THSLIEDERK KHYEGVMNSI ESFAKRTQIR SLQNYGTLTR TDLSNIYDRY HEVLSSKHRV 540
541 GLGSSTDTRL EFDEFCIFLA GVTEWAKGLD AAAINNSSSF LRHLFLRFDK SMTGSLSLQD 600
601 LVSGIAELKF RDVMRNISFI FELYDFNGDG FMDKPDVLKV SEAILWLTRF MGDEYLSAVS 660
661 EFIQRCFHFA DEASPDGHSD TLIDISDHMS STGSENRSVG ANSDIKVSLP TFRMVVLSIG 720
721 LLEQLFSGGL ADSIVLAPVQ EKSSTTGGLR GLLDSLVIDT NRIGKTFRGH KPASRPSTAN 780
781 GTSNQNTTSE ITTSETTATE KTPSNSSDTE DDVGDVVEND KDLLQFD |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
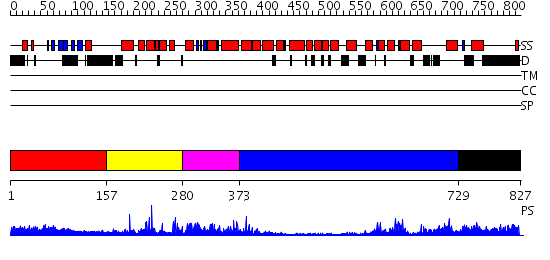
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..156] | 1.22 | No description for 2cayA was found. | |
| 2 | View Details | [157..279] | 46.39794 | No description for 2qq8A was found. | |
| 3 | View Details | [280..372] | 4.69897 | No description for 2jc6A was found. | |
| 4 | View Details | [373..728] | 5.39794 | Crystal Structure analysis of rat m-calpain mutant Lys10 Thr | |
| 5 | View Details | [729..827] | N/A | No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||
| 2 | No functions predicted. | |||||||||||||||||||||
| 3 | No functions predicted. | |||||||||||||||||||||
| 4 |
|
|||||||||||||||||||||
| 5 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.44 |
Source: Reynolds et al. (2008)