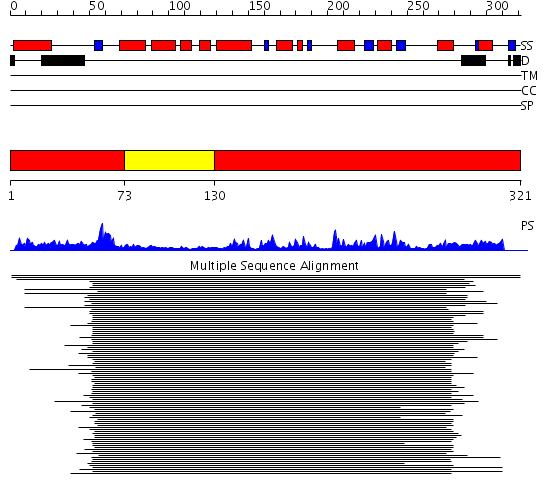
Description(s) found:
SHOW ONLY BEST
|
gi|731454|sp|P40025.1|PHM8_YEAST RecName: Full=Phosphate metabolism protein 8
[NCBI NR]
gi|6320875|ref|NP_010954.1| Protein of unknown function, expression is induced by low phosphate levels and by inactivation of Pho85p; Phm8p [Saccharomyces cerevisiae]
[NCBI NR]
gi|603270|gb|AAB64572.1| Yer037wp [Saccharomyces cerevisiae]
[NCBI NR]
pir||S50540 hypothetical protein YER037w - yeast (Saccharomyces cerevisiae)
[NCBI NR]
(P40025) Hypothetical 37.7 kDa protein in PIP1-GLN3 intergenic region
[Swiss-Prot]
Phosphate metabolism protein 8 OS=Saccharomyces cerevisiae GN=PHM8 PE=1 SV=1
[Swiss-Prot]
PHM8 Protein of unknown function, expression is induced by low phosphate levels and by inactivation of Pho85p
[mips]
Protein of unknown function, expression is induced by low phosphate levels and by inactivation of Pho85p
[SGD]
PHM8, Chr V from 225888-226853
[yeast_orfs2]
Chr V from 225888-226853
[yeastorf3.fasta]
PHM8 SGDID:S0000839, Chr V from 225888-226853, Verified ORF
[yeastflysequest.fasta]
Phm8p [Saccharomyces cerevisiae]�gi|731454|sp|P40025|YEM7_YEAST Hypothetical 37.7 kDa protein in PIP1-GLN3 intergenic region�gi|603270|gb|AAB64572.1| Yer037wp [Saccharomyces cerevisiae]�gi|1077647|pir||S50540 hypothetical protein YER037w - yeast (Saccharo
[nrdb_11152004_Chlamydomonas]
PHM8 SGDID:S000000839, Chr V from 225888-226853, Verified ORF, "Protein of unknown function, expression is induced by low phosphate levels and by inactivation of Pho85p"
[SGD_S-cerevisiae_na_12-16-2005_con_reversed.fasta]
Phosphate metabolism protein 8
[nr-271106-contam.fasta]
Yer037wp [Saccharomyces cerevisiae]
[nr-271106-contam.fasta]
Protein of unknown function, expression is induced by low phosphate levels and by inactivation of Pho85p; Phm8p [Saccharomyces cerevisiae]
[nr-271106-contam.fasta]
[chemostatdb.fasta]
Phosphate metabolism protein 8 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) GN=PHM8 PE=1 SV=1
[UniProt_S_cerevisiae_20120516_05-16-2012_reversed.fasta]
PHM8 SGDID:S000000839, Chr V from 225889-226854, Genome Release 64-1-1, Verified ORF, "Lysophosphatidic acid (LPA) phosphatase involved in LPA hydrolysis in response to phosphate starvation; phosphatase activity is soluble and Mg2+ dependent; expression is induced by low phosphate levels and by inactivation of Pho85p"
[yeast-030211-contam.fasta]
PHM8 SGDID:S000000839, Chr V from 225889-226854, Genome Release 64-2-1, Verified ORF, "Lysophosphatidic acid (LPA) phosphatase, nucleotidase; principle and physiological nucleotidase working on GMP, UMP and CMP; involved in LPA hydrolysis in response to phosphate starvation and ribose salvage pathway; phosphatase activity is soluble and Mg2+ dependent; expression is induced by low phosphate levels and by inactivation of Pho85p; PHM8 has a paralog, SDT1, that arose from the whole genome duplication"
[20141208_orf_trans.fasta]
TR4981|c0_g1_i1|g.3321 ORF TR4981|c0_g1_i1|g.3321 TR4981|c0_g1_i1|m.3321 type:complete len:322 (+) TR4981|c0_g1_i1:84-1049(+)
[DBVPG6044_S_cerevisiae-Sep2015-contam.fasta]
YER037W SGDID:S000000839, Chr V from 225889-226854, Genome Release 64-1-1, Verified ORF, "Lysophosphatidic acid (LPA) phosphatase involved in LPA hydrolysis in response to phosphate starvation; phosphatase activity is soluble and Mg2+ dependent; expression is induced by low phosphate levels and by inactivation of Pho85p"
[2016-01-SGD_Yeast_Contam_Riffle_20140507-Ska-Hec-110-tubulin.fasta]
asmbl_7542|g.5557 ORF asmbl_7542|g.5557 asmbl_7542|m.5557 type:complete len:322 (+) asmbl_7542:82-1047(+)
[DBVPG6044_Trinity_v2_2_0_PASA_6May2016-contam.fasta]
ID=asmbl_7520|g.4720 ORF ID=asmbl_7520|g.4720 ID=asmbl_7520|m.4720 type:complete len:322 (+) ID=asmbl_7520:82-1047(+)
[DBVPG6044_quiverGenomes_Trinity_v2_2_0_PASA_2June2016-contam.fasta]
|