













| General Information: |
|
| Name(s) found: |
lig3-PA /
FBpp0082071
[FlyBase]
|
| Description(s) found:
Found 15 descriptions. SHOW ALL |
|
| Organism: | Drosophila melanogaster |
| Length: | 806 amino acids |
Gene Ontology: |
|
| Cellular Component: |
intracellular
[IEA]
|
| Biological Process: |
DNA repair
[IEA]
DNA replication [IEA] DNA recombination [IEA] |
| Molecular Function: |
DNA binding
[IEA]
ATP binding [IEA] DNA ligase (ATP) activity [ISS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MLLSRLTWQA RILTRLLCNK MASKGLHNED NKLDTFRRIC DEIAGESSYL KKAERLQRFF 60
61 EKGSSGKGFK GDTVLWVQFL IPAANQRVYN LQNKALIKLF SRIFNADQQQ MHLDLEQDGD 120
121 VSETLRKHFA ASKKLKPQKQ SKLYLQEVEE MLLHLVERTK EDEQTDLLQK LCSKATDLDL 180
181 RTFIRLVKHD LRINARARHI LDAFGPDAYP AYQSSRDLVA IVEQFATKEN KNYASSPVKK 240
241 TKKTSSSAIQ VMTPISPMLA SPCKSVEDAF KKSPTGLYSE IKYDGERVQI HKQGNDFKFF 300
301 SRNLKPVVDH KIRALKKYIP QAFPGGGEMI LDSEIILVDT DTGALLPFGS LGAHKKQTFA 360
361 NAAVCLFVFD CILYDGEDLT QLPFRKRREI LEQNIQPIKS HVQLSESEFL KTTRELAAMT 420
421 ARVLHANLEG VVLKSPTSTY QPGKRNWLKV KKDYLFDGKM ADTADLVVLG AWYGSGKKGG 480
481 TLSIFLMGCY DAFGRLWKTV TKVHSGLDDA TNAEVHKSLI KLTERADANA IPSWLLCSKS 540
541 LVPDVLAKDP MKMPVWEITG AEFTKSDAHT ASGISVRFPR ITRRRSDKSA KEANDLAHLE 600
601 DLFEASKSNV NVDLLLANCS KDETAMNNKT TPEKEKKLNG NQSNGSKKLK RSNTGSDDED 660
661 GAPVLSKRKE SKAIRSLSPS QKGLKDFFKP SKEFKSVVKK EPKEDEGNIE STSSKPQLFE 720
721 GLVAHFAMEK DSSKLMQQFS QHGGCCTTEP QKADLVFYAN TNEALDLEKL RTLYRRSCRH 780
781 LNVSWLQSCL DTQSRQPFPL HAVMMK |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
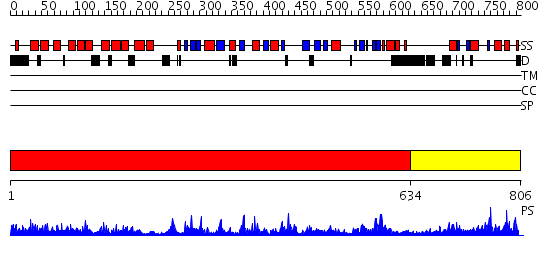
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..633] | 144.0 | Crystal Structure of Human DNA Ligase I bound to 5'-adenylated, nicked DNA | |
| 2 | View Details | [634..806] | 27.30103 | No description for 2faoA was found. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||
| 1 |
|
|||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.95 |
Source: Reynolds et al. (2008)